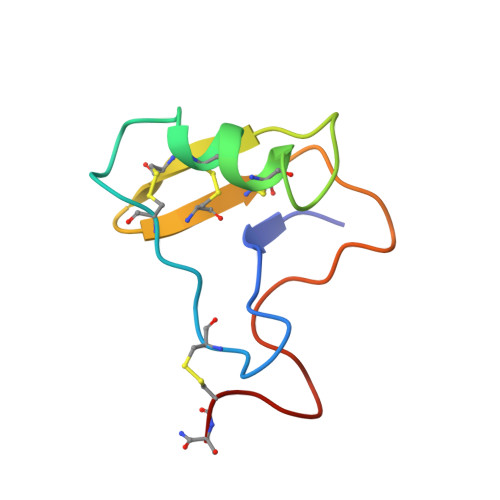
Solution structure of toxin 2 from centruroides noxius Hoffmann, a beta-scorpion neurotoxin acting on sodium channels.
Pintar, A., Possani, L.D., Delepierre, M.(1999) J Mol Biology 287: 359-367
- PubMed: 10080898 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2611
- Primary Citation Related Structures:
1CN2 - PubMed Abstract:
We have determined the solution structure of Cn2, a beta-toxin extracted from the venom of the New World scorpion Centruroides noxius Hoffmann. Cn2 belongs to the family of scorpion toxins that affect the sodium channel activity, and is very toxic to mammals (LD50=0.4 microg/20 g mouse mass). The three-dimensional structure was determined using 1H-1H two-dimensional NMR spectroscopy, torsion angle dynamics, and restrained energy minimization. The final set of 15 structures was calculated from 876 experimental distance constraints and 58 angle constraints. The structures have a global r. m.s.d. of 1.38 A for backbone atoms and 2.21 A for all heavy atoms. The overall fold is similar to that found in the other scorpion toxins acting on sodium channels. It is made of a triple-stranded antiparallel beta-sheet and an alpha-helix, and is stabilized by four disulfide bridges. A cis-proline residue at position 59 induces a kink of the polypeptide chain in the C-terminal region. The hydrophobic core of the protein is made up of residues L5, V6, L51, A55, and by the eight cysteine residues. A hydrophobic patch is defined by the aromatic residues Y4, Y40, Y42, W47 and by V57 on the side of the beta-sheet facing the solvent. A positively charged patch is formed by K8 and K63 on one edge of the molecule in the C-terminal region. Another positively charged spot is represented by the highly exposed K35. The structure of Cn2 is compared with those of other scorpion toxins acting on sodium channels, in particular Aah II and CsE-v3. This is the first structural report of an anti-mammal beta-scorpion toxin and it provides the necessary information for the design of recombinant mutants that can be used to probe structure-function relationships in scorpion toxins affecting sodium channel activity.
- Nuclear Magnetic Resonance Laboratory, URA1129 Aids-Retrovirus Department, Pasteur Institute, 28 Rue du Dr Roux, Paris, 75015, France.
Organizational Affiliation: