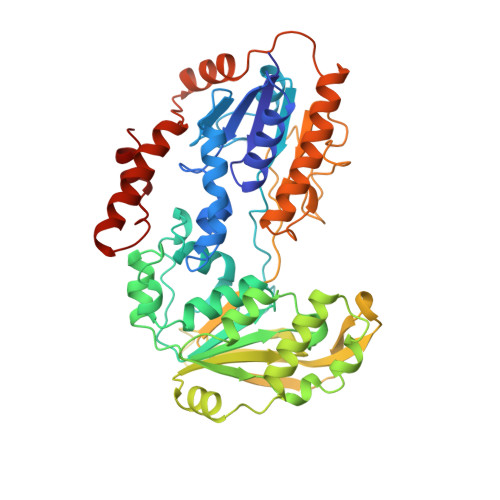
The structure of adrenodoxin reductase of mitochondrial P450 systems: electron transfer for steroid biosynthesis.
Ziegler, G.A., Vonrhein, C., Hanukoglu, I., Schulz, G.E.(1999) J Mol Biology 289: 981-990
- PubMed: 10369776 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2807
- Primary Citation Related Structures:
1CJC - PubMed Abstract:
Adrenodoxin reductase is a monomeric 51 kDa flavoenzyme that is involved in the biosynthesis of all steroid hormones. The structure of the native bovine enzyme was determined at 2.8 A resolution, and the structure of the respective recombinant enzyme at 1.7 A resolution. Adrenodoxin reductase receives a two-electron package from NADPH and converts it to two single electrons that are transferred via adrenodoxin to all mitochondrial cytochromes P 450. The structure suggests how the observed flavin semiquinone is stabilized. A striking feature is the asymmetric charge distribution, which most likely controls the approach of the electron carrier adrenodoxin. A model for the interaction is proposed. Adrenodoxin reductase shows clear sequence homology to half a dozen proteins identified in genome analysis projects, but neither sequence nor structural homology to established, functionally related electron transferases. Yet, the structure revealed a relationship to the disulfide oxidoreductases, permitting the assignment of the NADP-binding site.
- Albert-Ludwigs-Universität, Albertstrasse 21, Freiburg im Breisgau, D-79104, Germany.
Organizational Affiliation: