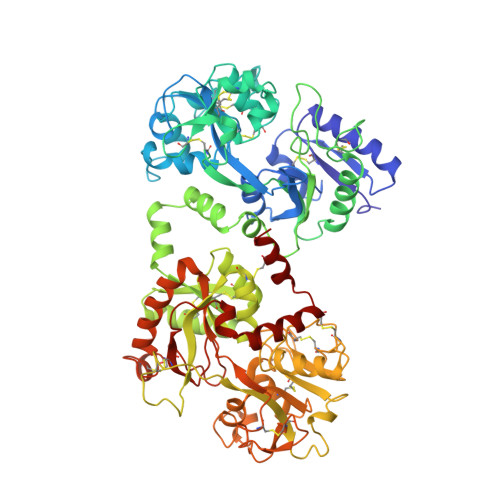
Structure of buffalo lactoferrin at 2.5 A resolution using crystals grown at 303 K shows different orientations of the N and C lobes.
Karthikeyan, S., Paramasivam, M., Yadav, S., Srinivasan, A., Singh, T.P.(1999) Acta Crystallogr D Biol Crystallogr 55: 1805-1813
- PubMed: 10531476 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444999010951
- Primary Citation Related Structures:
1CE2 - PubMed Abstract:
The structure of buffalo lactoferrin has been determined at 303 K. The crystals belong to orthorhombic space group P2(1)2(1)2(1), with unit-cell parameters a = 77.5, b = 91.0, c = 131.5 A and Z = 4. The structure has been refined to an R factor of 0.187. The overall structure of the protein is similar to its structure determined at 277 K in a different crystal form. However, the lobe orientations in the two structures differ by 9.0 degrees, suggesting significant inter-lobe flexibility in this family of proteins. The inter-lobe interactions are predominantly hydrophobic and could act as a cushion for a change in orientation under the influence of external conditions. On the other hand, the domain arrangements are found to be similar in 277 and 303 K crystal structures, with orientations differing by 1.5 and 1.0 degrees in the N and C lobes, respectively. The results of these investigations suggest that the increase in temperature helps in the production of better quality crystals.
- Department of Biophysics, All India Institute of Medical Sciences, New Delhi 110 029, India.
Organizational Affiliation: