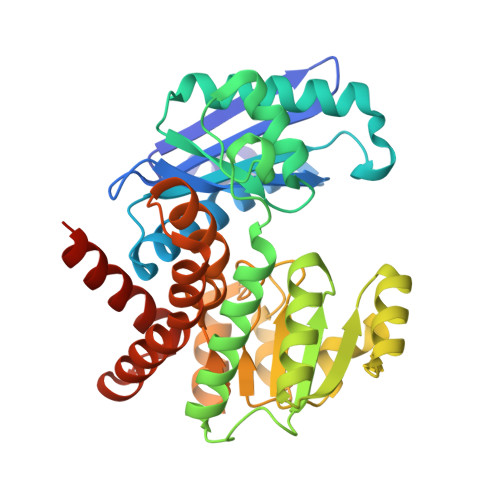
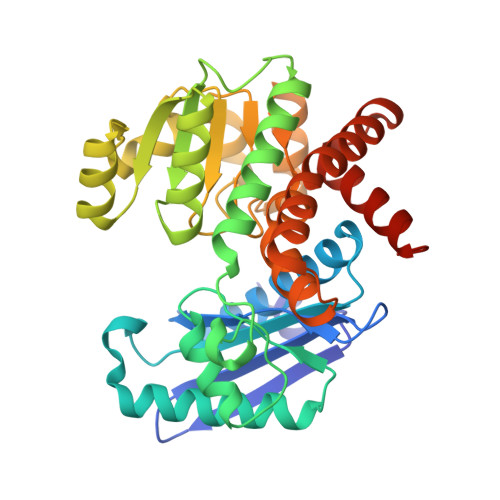
Rhodococcus L-phenylalanine dehydrogenase: kinetics, mechanism, and structural basis for catalytic specificity.
Brunhuber, N.M., Thoden, J.B., Blanchard, J.S., Vanhooke, J.L.(2000) Biochemistry 39: 9174-9187
- PubMed: 10924111 Search on PubMed
- DOI: https://doi.org/10.1021/bi000494c
- Primary Citation Related Structures:
1C1D, 1C1X - PubMed Abstract:
Phenylalanine dehydrogenase catalyzes the reversible, pyridine nucleotide-dependent oxidative deamination of L-phenylalanine to form phenylpyruvate and ammonia. We have characterized the steady-state kinetic behavior of the enzyme from Rhodococcus sp. M4 and determined the X-ray crystal structures of the recombinant enzyme in the complexes, E.NADH.L-phenylalanine and E.NAD(+). L-3-phenyllactate, to 1.25 and 1.4 A resolution, respectively. Initial velocity, product inhibition, and dead-end inhibition studies indicate the kinetic mechanism is ordered, with NAD(+) binding prior to phenylalanine and the products' being released in the order of ammonia, phenylpyruvate, and NADH. The enzyme shows no activity with NADPH or other 2'-phosphorylated pyridine nucleotides but has broad activity with NADH analogues. Our initial structural analyses of the E.NAD(+).phenylpyruvate and E.NAD(+). 3-phenylpropionate complexes established that Lys78 and Asp118 function as the catalytic residues in the active site [Vanhooke et al. (1999) Biochemistry 38, 2326-2339]. We have studied the ionization behavior of these residues in steady-state turnover and use these findings in conjunction with the structural data described both here and in our first report to modify our previously proposed mechanism for the enzymatic reaction. The structural characterizations also illuminate the mechanism of the redox specificity that precludes alpha-amino acid dehydrogenases from functioning as alpha-hydroxy acid dehydrogenases.
- Department of Biochemistry, Albert Einstein College of Medicine, Bronx, New York 10461, USA.
Organizational Affiliation: