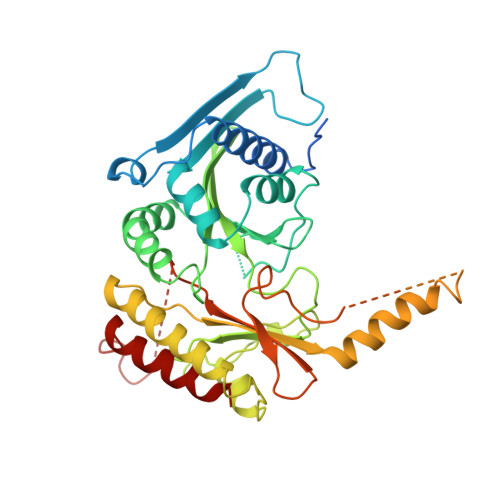
Structure of type IIbeta phosphatidylinositol phosphate kinase: a protein kinase fold flattened for interfacial phosphorylation.
Rao, V.D., Misra, S., Boronenkov, I.V., Anderson, R.A., Hurley, J.H.(1998) Cell 94: 829-839
- PubMed: 9753329 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(00)81741-9
- Primary Citation Related Structures:
1BO1 - PubMed Abstract:
Phosphoinositide kinases play central roles in signal transduction by phosphorylating the inositol ring at specific positions. The structure of one such enzyme, type IIbeta phosphatidylinositol phosphate kinase, reveals a protein kinase ATP-binding core and demonstrates that all phosphoinositide kinases belong to one superfamily. The enzyme is a disc-shaped homodimer with a 33 x 48 A basic flat face that suggests an electrostatic mechanism for plasma membrane targeting. Conserved basic residues form a putative phosphatidylinositol phosphate specificity site. The substrate-binding site is open on one side, consistent with dual specificity for phosphatidylinositol 3- and 5-phosphates. A modeled complex with membrane-bound substrate and ATP shows how a phosphoinositide kinase can phosphorylate its substrate in situ at the membrane interface.
- Laboratory of Molecular Biology, National Institute of Diabetes, Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892-0580, USA.
Organizational Affiliation: