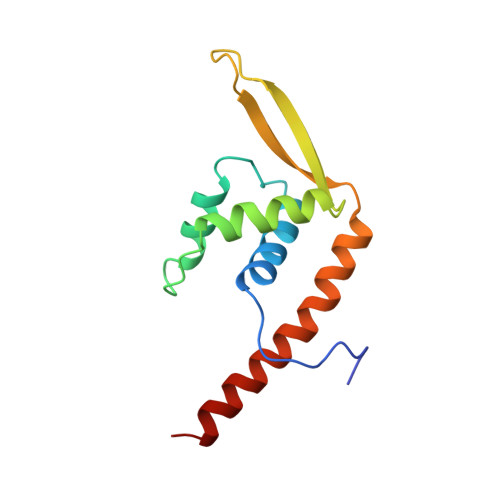
Crystal structure of the replication terminator protein from B. subtilis at 2.6 A.
Bussiere, D.E., Bastia, D., White, S.W.(1995) Cell 80: 651-660
- PubMed: 7867072 Search on PubMed
- DOI: https://doi.org/10.1016/0092-8674(95)90519-7
- Primary Citation Related Structures:
1BM9 - PubMed Abstract:
The crystal structure of the replication terminator protein (RTP) of B. subtilis has been determined at 2.6 A resolution. As previously suggested by both biochemical and biophysical studies, the molecule exists as a symmetric dimer and is in the alpha + beta protein-folding class. The protein has several uncommon features, including an antiparallel coiled-coil, which serves as the dimerization domain, and both an alpha-helix and a beta-ribbon suitably positioned to interact with the major and minor grooves of B-DNA. A site has been identified on the surface of RTP that is biochemically and positionally suitable for interaction with the replication-specific helicase. Other features of the structure are consistent with the polar contrahelicase mechanism of the protein. A model of the interaction between RTP and its cognate DNA is presented.
- Department of Microbiology, Duke University Medical Center, Durham, North Carolina 27710.
Organizational Affiliation: