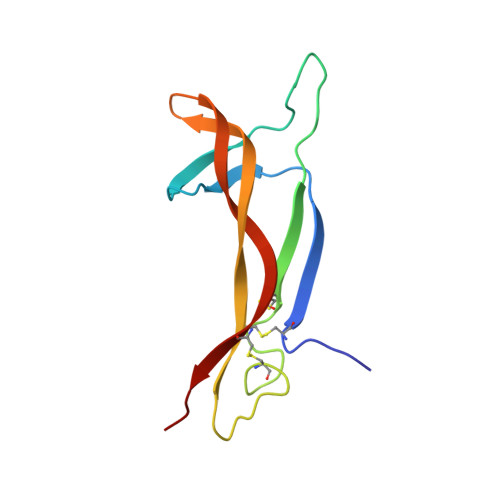
New protein fold revealed by a 2.3-A resolution crystal structure of nerve growth factor.
McDonald, N.Q., Lapatto, R., Murray-Rust, J., Gunning, J., Wlodawer, A., Blundell, T.L.(1991) Nature 354: 411-414
- PubMed: 1956407 Search on PubMed
- DOI: https://doi.org/10.1038/354411a0
- Primary Citation Related Structures:
1BET - PubMed Abstract:
Nerve growth factor (NGF) is a member of an expanding family of neurotrophic factors (including brain-derived neurotrophic factor and the neurotrophins) that control the development and survival of certain neuronal populations both in the peripheral and in the central nervous systems. Its biological effects are mediated by a high-affinity ligand-receptor interaction and a tyrosine kinase signalling pathway. A potential use for NGF and its relatives in the treatment of neurological disorders such as Alzheimer's disease and Parkinson's disease requires an understanding of the structure-function relationships of NGF. NGF is a dimeric molecule, with 118 amino acids per protomer. We report the crystal structure of the murine NGF dimer at 2.3-A resolution, which reveals a novel protomer structure consisting of three antiparallel pairs of beta strands, together forming a flat surface. Two subunits associate through this surface, thus burying a total of 2,332 A. Four loop regions, which contain many of the variable residues observed between different NGF-related molecules, may determine the different receptor specificities. A clustering of positively charged side chains may provide a complementary interaction with the acidic low-affinity NGF receptor. The structure provides a model for rational design of analogues of NGF and its relatives and for testing the NGF-receptor recognition determinants critical for signal transduction.
- ICRF Unit for Structural Molecular Biology, Birkbeck College, London, UK.
Organizational Affiliation: