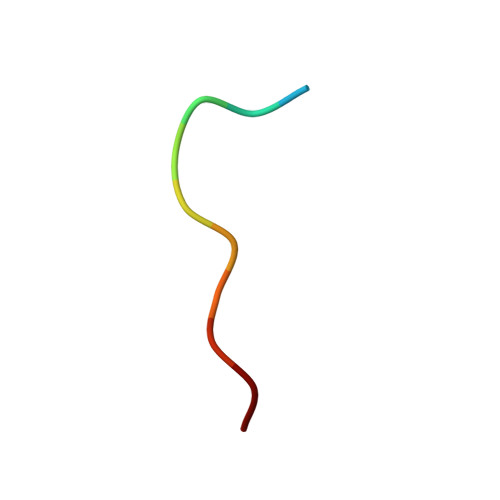
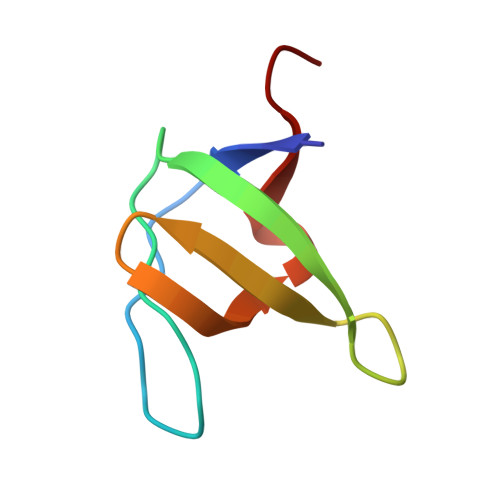
Crystal structure of the abl-SH3 domain complexed with a designed high-affinity peptide ligand: implications for SH3-ligand interactions.
Pisabarro, M.T., Serrano, L., Wilmanns, M.(1998) J Mol Biology 281: 513-521
- PubMed: 9698566 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.1932
- Primary Citation Related Structures:
1BBZ - PubMed Abstract:
The Abl-SH3 domain is implicated in negative regulation of the Abl kinase by mediating protein-protein interactions. High-affinity SH3 ligands could compete for these interactions and specifically activate the Abl kinase, providing control and a better understanding of the molecular interactions that underlie diseases where SH3 domains are involved. The p41 peptide (APSYSPPPPP) is a member of a group of peptide ligands designed to bind specifically the Abl-SH3 domain. It binds to Abl-SH3 with a Kd of 1.5 microM, whereas its affinity for the Fyn-SH3 domain is 273 microM. We have determined the crystal structure of the Abl-SH3 domain in complex with the high-affinity peptide ligand p41 at 1.6 A resolution. In the crystal structure, this peptide adopts a polyproline type II helix conformation through residue 5 to 10, and it binds in type I orientation to the Abl-SH3 domain. The tyrosine side-chain in position 4 of the peptide is hydrogen bonded to two residues in the RT-loop of the Abl-SH3 domain. The tight fit of this side-chain into the RT-loop pocket is enhanced by conformational adjustment of the main chain at position 5. The SH3 ligand peptides can be divided into two distinct parts. The N-terminal part binds to the SH3 domain in the region formed by the valley between the nSrc and RT-loops. It determines the specificity for different SH3 domains. The C-terminal part adopts a polyproline type II helix conformation. This binds to a well-conserved hydrophobic surface of the SH3 domain. Analysis of two "half"-peptides, corresponding to these ligand parts, shows that both are essential components for strong binding to the SH3 domains. The crystal structure of the Abl-SH3:p41 complex explains the high affinity and specificity of the p41 peptide towards the Abl-SH3 domain, and reveals principles that will be exploited for future design of small, high-affinity ligands to interfere efficiently with the in vivo regulation of Abl kinase activity.
- EMBL, Structures & Biocomputing, Meyerhofstrasse 1, Heidelberg, 69117, Germany. pisabarro@embl-heidelberg.de
Organizational Affiliation: