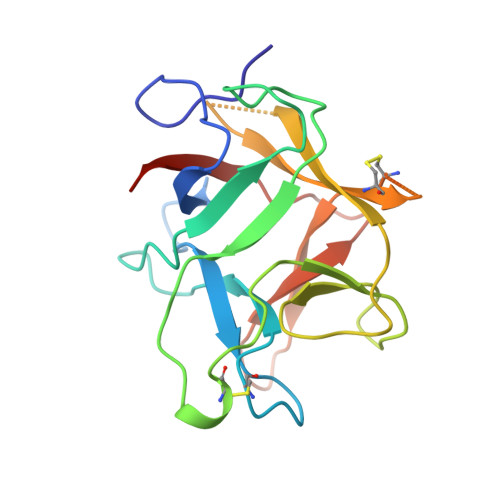
Structure of the Kunitz-type soybean trypsin inhibitor (STI): implication for the interactions between members of the STI family and tissue-plasminogen activator.
De Meester, P., Brick, P., Lloyd, L.F., Blow, D.M., Onesti, S.(1998) Acta Crystallogr D Biol Crystallogr 54: 589-597
- PubMed: 9761854 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444997015849
- Primary Citation Related Structures:
1BA7 - PubMed Abstract:
The Kunitz-type soybean trypsin inhibitor (STI) has played a key role in the early study of proteinases, having been used as the main substrate in the biochemical and kinetic work that led to the definition of the standard mechanism of action of proteinase inhibitors. A partial structure of STI complexed with porcine trypsin has previously been reported, in which the first 93 residues of the inhibitor, including the region of contact with trypsin, were relatively well defined, whereas for the remaining part of the peptide chain only some Calpha atoms were located. The structure of the inhibitor in its free form has now been determined by molecular replacement to 2.5 A, using the coordinates of the homologous Erythrina trypsin inhibitor as a search model. When the refined atomic coordinates of STI are compared with the partial model previously available, the conformation of the reactive-site loop and its position with respect to the main body of the molecule does not change when the inhibitor interacts with trypsin. There are instead, despite the high similarity in the overall tertiary structure, significant differences between STI and Erythrina trypsin inhibitor (ETI) in the region which is in contact with the enzyme in the STI:trypsin crystal structure. Some of these differences can explain the unique specificity of ETI and its ability to inhibit the fibrinolytic enzyme tissue-type plasminogen activator.
- Blackett Laboratory, Imperial College, London SW7 2BZ, England.
Organizational Affiliation: