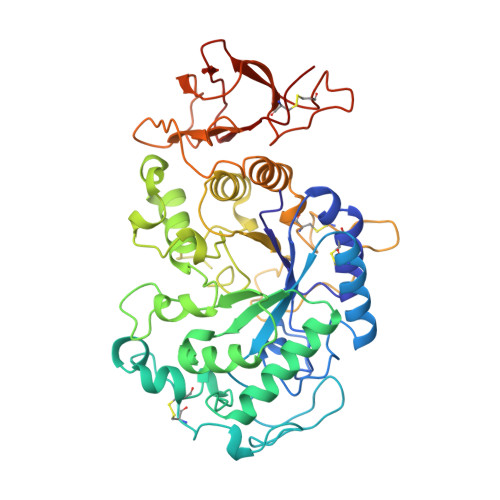
Structures of the psychrophilic Alteromonas haloplanctis alpha-amylase give insights into cold adaptation at a molecular level.
Aghajari, N., Feller, G., Gerday, C., Haser, R.(1998) Structure 6: 1503-1506
- PubMed: 9862804 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(98)00149-x
- Primary Citation Related Structures:
1B0I - PubMed Abstract:
. Enzymes from psychrophilic (cold-adapted) microorganisms operate at temperatures close to 0 degreesC, where the activity of their mesophilic and thermophilic counterparts is drastically reduced. It has generally been assumed that thermophily is associated with rigid proteins, whereas psychrophilic enzymes have a tendency to be more flexible. . Insights into the cold adaptation of proteins are gained on the basis of a psychrophilic protein's molecular structure. To this end, we have determined the structure of the recombinant form of a psychrophilic alpha-amylase from Alteromonas haloplanctis at 2.4 A resolution. We have compared this with the structure of the wild-type enzyme, recently solved at 2.0 A resolution, and with available structures of their mesophilic counterparts. These comparative studies have enabled us to identify possible determinants of cold adaptation. . We propose that an increased resilience of the molecular surface and a less rigid protein core, with less interdomain interactions, are determining factors of the conformational flexibility that allows efficient enzyme catalysis in cold environments.
- Institut de Biologie et Chimie des Protéines UPR 412, CNRS 7 Passage du Vercors 69367 Lyon cedex 07 France.
Organizational Affiliation: