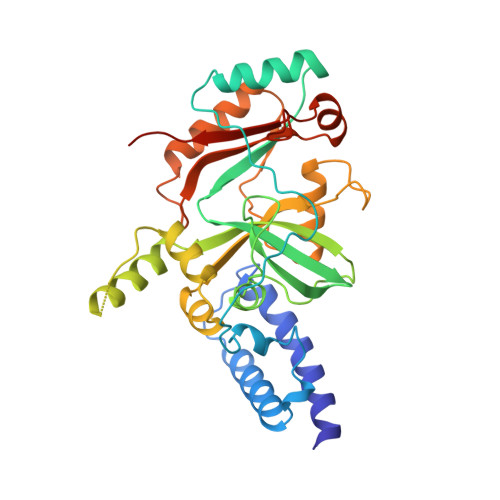
Structure of the adenylation domain of an NAD+-dependent DNA ligase.
Singleton, M.R., Hakansson, K., Timson, D.J., Wigley, D.B.(1999) Structure 7: 35-42
- PubMed: 10368271 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(99)80007-0
- Primary Citation Related Structures:
1B04 - PubMed Abstract:
DNA ligases catalyse phosphodiester bond formation between adjacent bases in nicked DNA, thereby sealing the nick. A key step in the catalytic mechanism is the formation of an adenylated DNA intermediate. The adenyl group is derived from either ATP (in eucaryotes and archaea) or NAD+4 (in bacteria). This difference in cofactor specificity suggests that DNA ligase may be a useful antibiotic target. The crystal structure of the adenylation domain of the NAD+-dependent DNA ligase from Bacillus stearothermophilus has been determined at 2.8 A resolution. Despite a complete lack of detectable sequence similarity, the fold of the central core of this domain shares homology with the equivalent region of ATP-dependent DNA ligases, providing strong evidence for the location of the NAD+-binding site. Comparison of the structure of the NAD+4-dependent DNA ligase with that of ATP-dependent ligases and mRNA-capping enzymes demonstrates the manifold utilisation of a conserved nucleotidyltransferase domain within this family of enzymes. Whilst this conserved core domain retains a common mode of nucleotide binding and activation, it is the additional domains at the N terminus and/or the C terminus that provide the alternative specificities and functionalities in the different members of this enzyme superfamily.
- Sir William Dunn School of Pathology, University of Oxford, South ParksRoad, Oxford OX1 3RE, UK.
Organizational Affiliation: