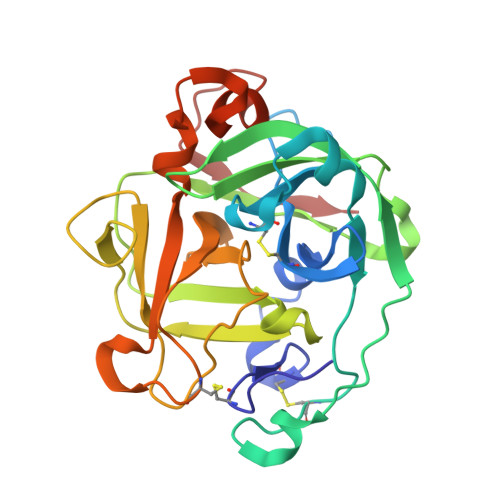
The primary structure and structural characteristics of Achromobacter lyticus protease I, a lysine-specific serine protease.
Tsunasawa, S., Masaki, T., Hirose, M., Soejima, M., Sakiyama, F.(1989) J Biological Chem 264: 3832-3839
- PubMed: 2492988 Search on PubMed
- Primary Citation Related Structures:
1ARB, 1ARC - PubMed Abstract:
The complete amino acid sequence of Achromobacter lyticus protease I (EC 3.4.21.50), which specifically hydrolyzes lysyl peptide bonds, has been established. This has been achieved by sequence analysis of the reduced and S-carboxymethylated protease and of peptides obtained by enzymatic digestion with Achromobacter protease I itself and Staphylococcus aureus V8 protease and by chemical cleavage with cyanogen bromide. The protease consists of 268 residues with three disulfide bonds, which have been assigned to Cys6-Cys216, Cys12-Cys80, and Cys36-Cys58. Comparison of the amino acid sequence of Achromobacter protease and other serine proteases of bacterial and mammalian origins has revealed that Achromobacter protease I is a mammalian-type serine protease of which the catalytic triad comprises His57, Asp113, and Ser194. It has also been shown that the protease has 9- and 26-residue extensions of the peptide chain at the N and C termini, respectively, and overall sequence homology is as low as 20% with bovine trypsin. The presence of a disulfide bridge between the N-terminal extension Cys6 and Cys216 close to the putative active site in the C-terminal region is thought to be responsible for the generation of maximal proteolytic function in the pH range 8.5-10.7 and enhanced stability to denaturation.
- Institute for Protein Research, Osaka University, Japan.
Organizational Affiliation: