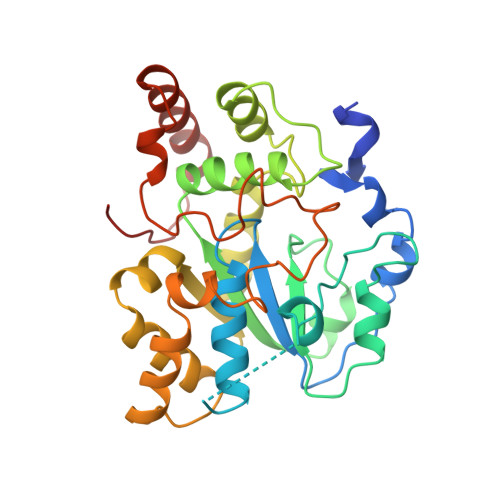
Crystal structure of estrogen sulphotransferase.
Kakuta, Y., Pedersen, L.G., Carter, C.W., Negishi, M., Pedersen, L.C.(1997) Nat Struct Biol 4: 904-908
- PubMed: 9360604 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1197-904
- Primary Citation Related Structures:
1AQU, 1AQY - PubMed Abstract:
The structure of estrogen sulphotransferase has been solved in the presence of inactive cofactor PAP and substrate 17 beta-estradiol. This structure reveals structural similarities between cytosolic sulphotransferases and nucleotide kinases.