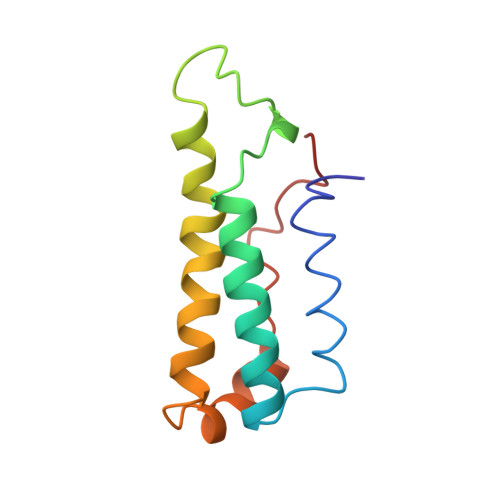
Solution structure of apocytochrome b562.
Feng, Y., Sligar, S.G., Wand, A.J.(1994) Nat Struct Biol 1: 30-35
- PubMed: 7656004 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0194-30
- Primary Citation Related Structures:
1APC - PubMed Abstract:
The apoprotein is an important intermediate on the folding pathways of many haem proteins, yet a detailed structure of such an intermediate has remained elusive. Here we present the structure of apocytochrome b562 obtained by NMR spectroscopy. The apoprotein has a topology similar to the holoprotein. Nevertheless, significant differences in helix-helix packing between the two are evident. Much of the haem binding pocket in the apoprotein is preserved but exposed to solvent creating a large cavern. As apocytochrome b562 displays many of the physical characteristics ascribed to the molten globule state, these results help ellucidate the origin of several properties of the protein molten globule.
- Department of Biochemistry, 415 Roger Adams Laboratory, University of Illinois at Urbana-Champaign 61801, USA.
Organizational Affiliation: