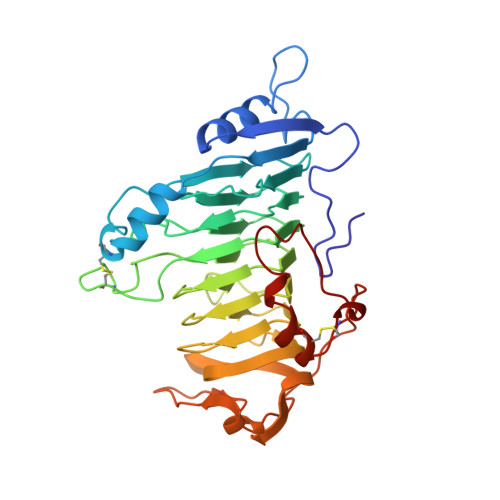
The Refined Three-Dimensional Structure of Pectate Lyase E from Erwinia chrysanthemi at 2.2 A Resolution.
Lietzke, S.E., Scavetta, R.D., Yoder, M.D., Jurnak, F.(1996) Plant Physiol 111: 73-92
- PubMed: 12226275 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1104/pp.111.1.73
- Primary Citation Related Structures:
1AIR - PubMed Abstract:
The crystal structure of pectate lyase E (PelE; EC 4.2.2.2) from the enterobacteria Erwinia chrysanthemi has been refined by molecular dynamics techniques to a resolution of 2.2 A and an R factor (an agreement factor between observed structure factor amplitudes) of 16.1%. The final model consists of all 355 amino acids and 157 water molecules. The root-mean-square deviation from ideality is 0.009 A for bond lengths and 1.721[deg] for bond angles. The structure of PelE bound to a lanthanum ion, which inhibits the enzymatic activity, has also been refined and compared to the metal-free protein. In addition, the structures of pectate lyase C (PelC) in the presence and absence of a lutetium ion have been refined further using an improved algorithm for identifying waters and other solvent molecules. The two putative active site regions of PelE have been compared to those in the refined structure of PelC. The analysis of the atomic details of PelE and PelC in the presence and absence of lanthanide ions provides insight into the enzymatic mechanism of pectate lyases.
- Department of Biochemistry, University of California, Riverside, California 92521.
Organizational Affiliation: