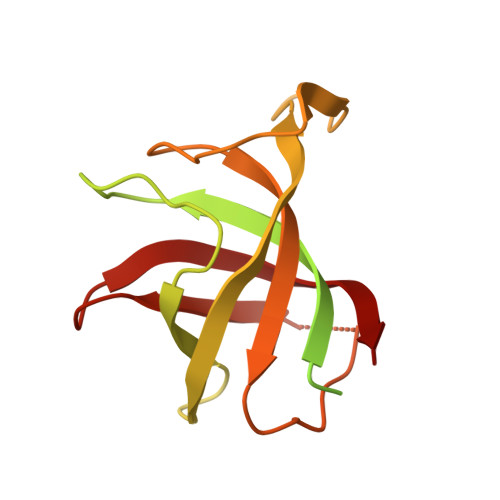
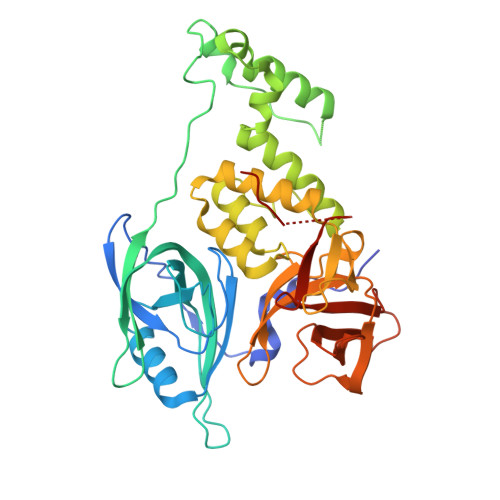
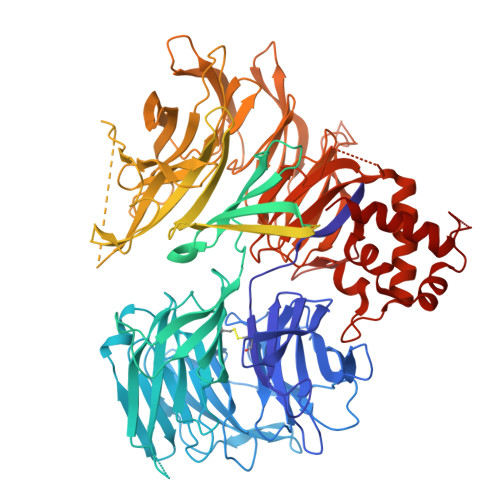
TNG961 is a selective oral HBS1L molecular glue degrader for the treatment of FOCAD-deleted cancers.
Nicholson, H.E., Whittington, D.A., Bruzzese, F.J., Lazarides, K., Martires, L.C.M., Tonini, M.R., Jenkins, H.N., Zhang, M., Shahagadkar, P., Pratt, C.B., Briggs, K.J., McCarren, P., Tsai, A., Bandi, M., Min, C., Huang, A., Zhang, H., Meier, S.R., Shen, B., Yu, Y., Liang, C., Liu, Y., Teng, T., Zhang, J., Crystal, A., Mallender, W.D., Wu, X.E., Maxwell, J.P., Andersen, J.N.(2026) Cancer Discov
- PubMed: 42001523 Search on PubMed
- DOI: https://doi.org/10.1158/2159-8290.CD-26-0040
- Primary Citation Related Structures:
10AY, 11MR - PubMed Abstract:
When tumor suppressor genes are lost through chromosomal deletion, deletion of adjacent genes can generate therapeutic vulnerabilities. MTAP is frequently co-deleted with the Chr9p21 tumor suppressor gene CDKN2A, creating synthetic lethal dependency on PRMT5. Telomeric to MTAP lies FOCAD, whose loss induces dependency on the HBS1L/PELO ribosome-rescue complex for translational maintenance. FOCAD is deleted in ~1/3 of MTAP-deleted cancers. We screened an IMiD-focused diversity library and identified a weak hit that bound cereblon, promoted HBS1L-CRBN-compound complex formation, and induced E3-ligase-dependent HBS1L ubiquitination and degradation. Guided by cryo-EM structures and proteome selectivity we developed TNG961, a potent, selective HBS1L degrader that disrupts the HBS1L/PELO complex, inducing translational arrest, unfolded protein response activation, and growth inhibition in FOCAD-negative models. Oral administration of TNG961 regresses FOCAD-negative xenografts, including PRMT5 inhibitor-refractory models, establishing HBS1L degradation as a strategy to exploit FOCAD loss and supporting clinical evaluation of TNG961 as a first-in-class precision oncology therapeutic.
- Tango Therapeutics (United States) Boston United States.
Organizational Affiliation: