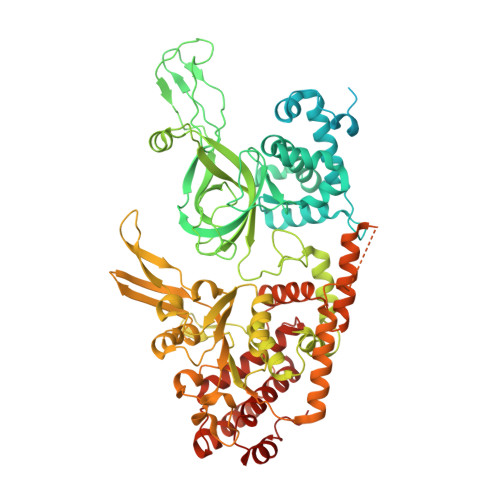
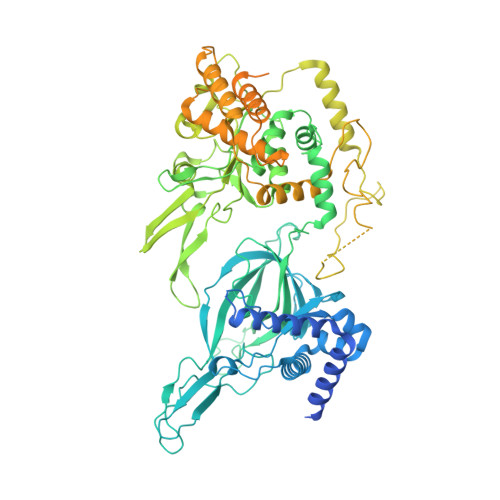
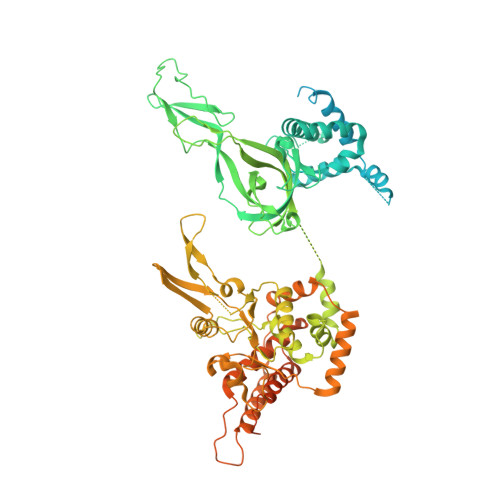
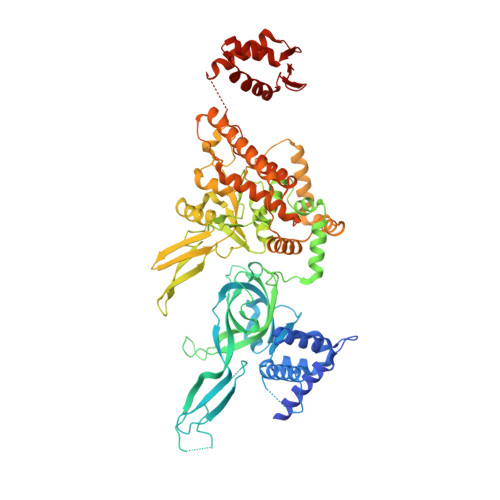
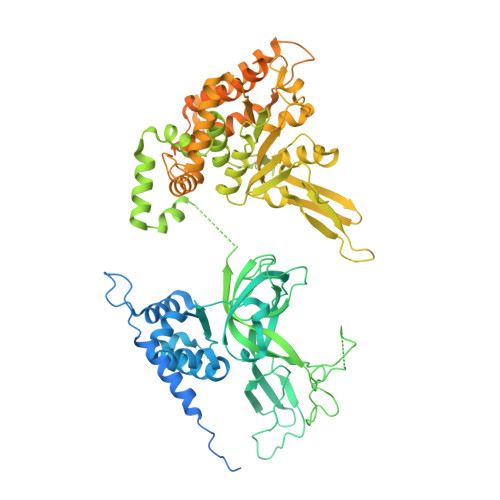
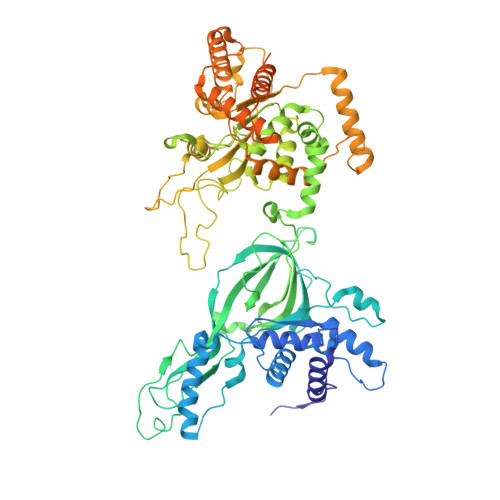
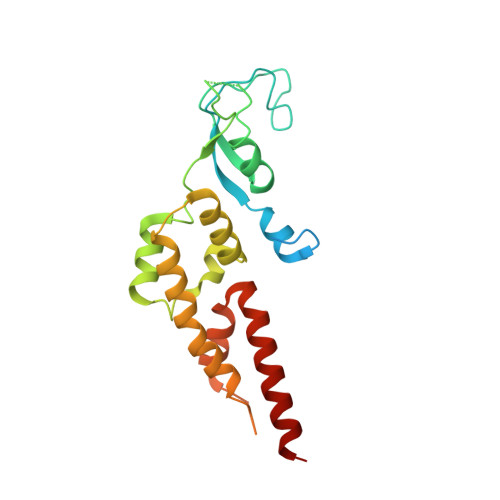
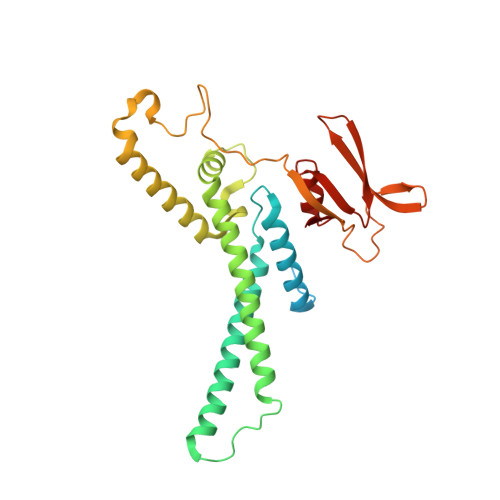
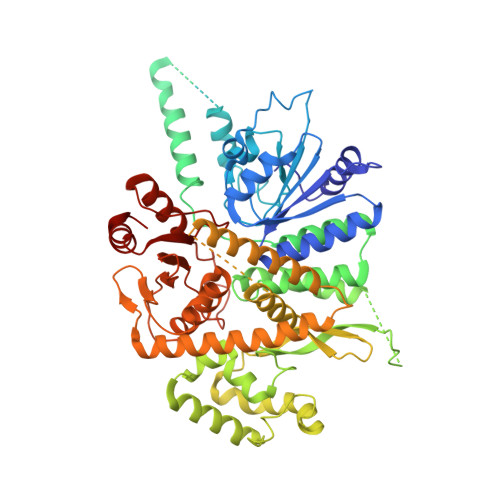
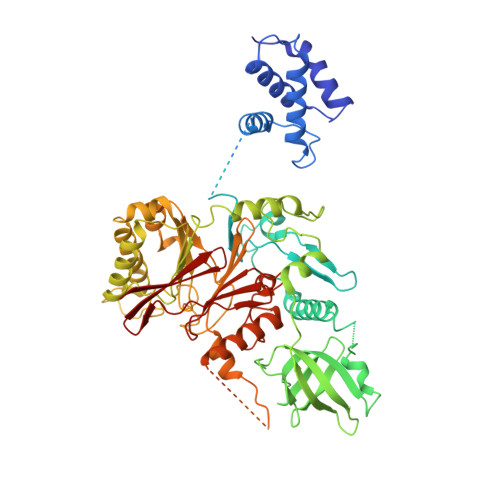
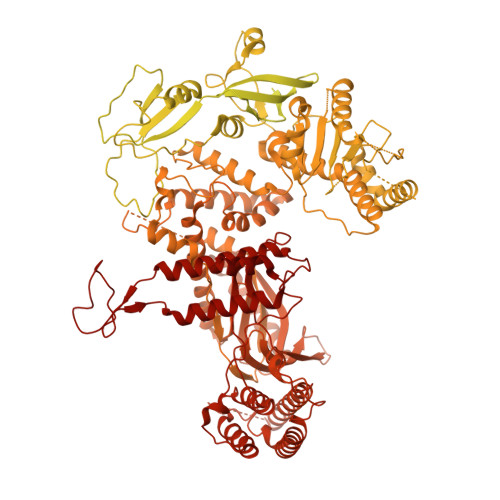
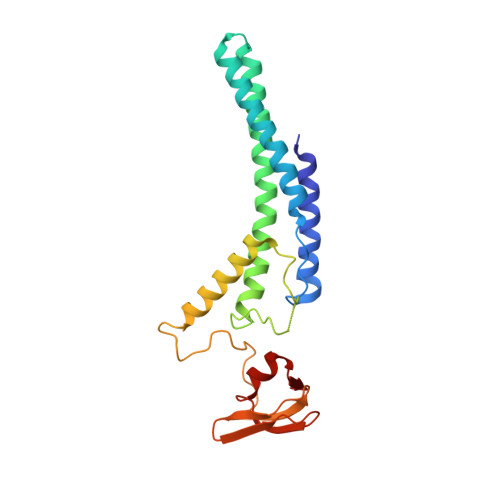
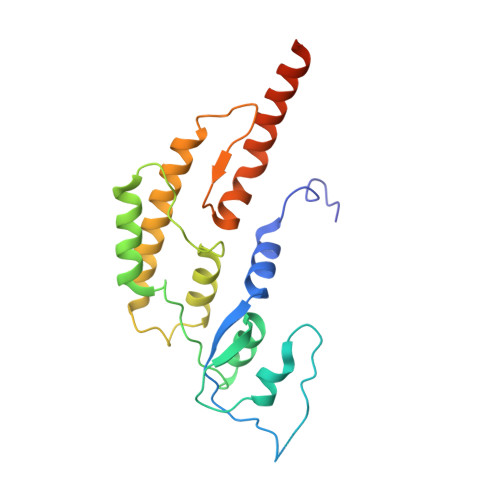
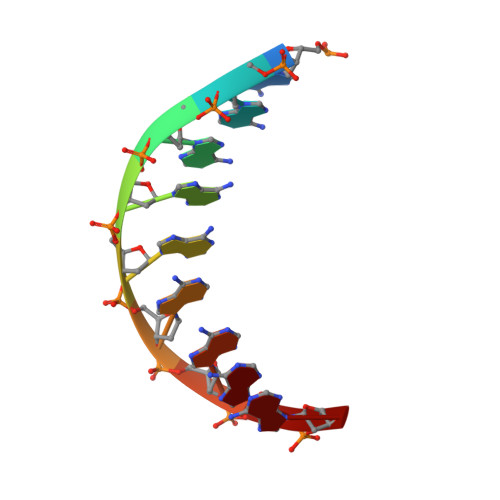
Unwinding of a eukaryotic origin of replication visualized by cryo-EM.
Henrikus, S.S., Gross, M.H., Willhoft, O., Puhringer, T., Lewis, J.S., McClure, A.W., Greiwe, J.F., Palm, G., Nans, A., Diffley, J.F.X., Costa, A.(2024) Nat Struct Mol Biol 31: 1265-1276
- PubMed: 38760633
- DOI: https://doi.org/10.1038/s41594-024-01280-z
- Primary Citation of Related Structures:
8P5E, 8P62, 8P63 - PubMed Abstract:
To prevent detrimental chromosome re-replication, DNA loading of a double hexamer of the minichromosome maintenance (MCM) replicative helicase is temporally separated from DNA unwinding. Upon S-phase transition in yeast, DNA unwinding is achieved in two steps: limited opening of the double helix and topological separation of the two DNA strands. First, Cdc45, GINS and Polε engage MCM to assemble a double CMGE with two partially separated hexamers that nucleate DNA melting. In the second step, triggered by Mcm10, two CMGEs separate completely, eject the lagging-strand template and cross paths. To understand Mcm10 during helicase activation, we used biochemical reconstitution with cryogenic electron microscopy. We found that Mcm10 splits the double CMGE by engaging the N-terminal homo-dimerization face of MCM. To eject the lagging strand, DNA unwinding is started from the N-terminal side of MCM while the hexamer channel becomes too narrow to harbor duplex DNA.
Organizational Affiliation:
Macromolecular Machines Laboratory, Francis Crick Institute, London, UK.