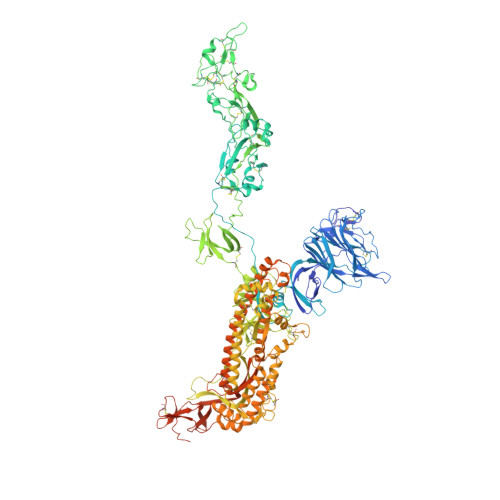
Structural basis for the recognition of HCoV-HKU1 by human TMPRSS2.
Xia, L., Zhang, Y., Zhou, Q.(2024) Cell Res
- PubMed: 38641728
- DOI: https://doi.org/10.1038/s41422-024-00958-9
- Primary Citation of Related Structures:
8Y19, 8Y1A, 8Y1B, 8Y1C, 8Y1D, 8Y1E, 8Y1F, 8Y1G, 8Y1H
Organizational Affiliation:
Center for Infectious Disease Research, Research Center for Industries of the Future, Zhejiang Key Laboratory of Structural Biology, School of Life Sciences, Westlake University. Institute of Biology, Westlake Institute for Advanced Study, Hangzhou, Zhejiang, China.