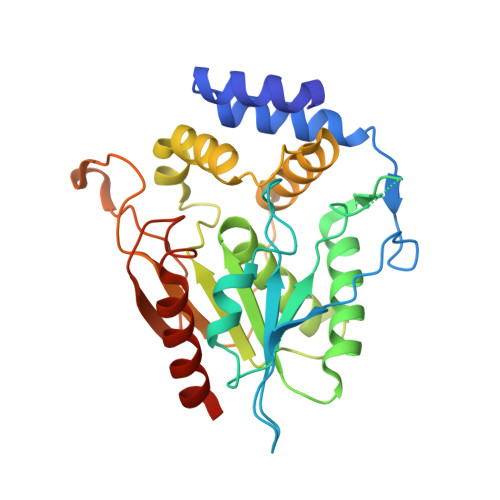
Substrate Trapping in Polyketide Synthase Thioesterase Domains: Structural Basis for Macrolactone Formation
McCullough, T.M., Choudhary, V., Akey, D.L., Skiba, M.A., Bernard, S.M., Kittendorf, J.D., Schmidt, J.J., Sherman, D.H., Smith, J.L.(2024) ACS Catal 14: 12551-12563