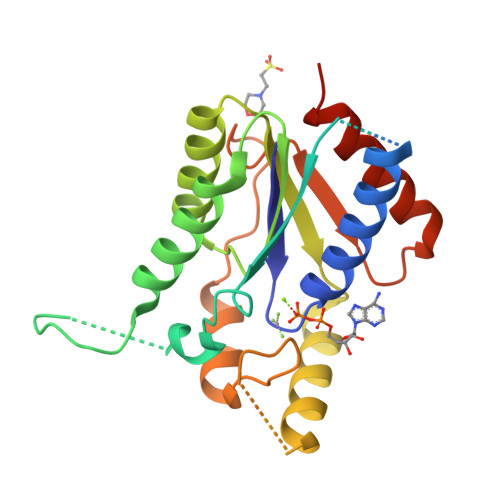
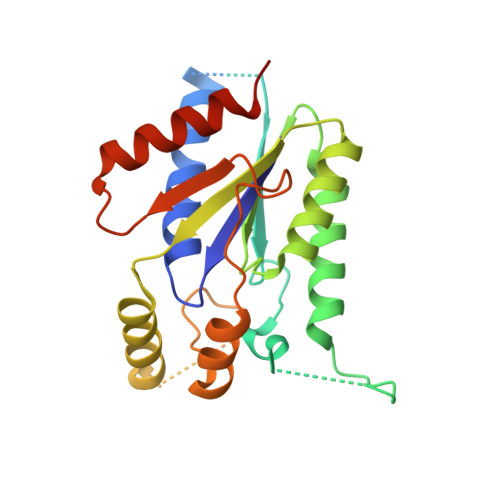
Kti12, a PSTK-like tRNA dependent ATPase essential for tRNA modification by Elongator.
Krutyholowa, R., Hammermeister, A., Zabel, R., Abdel-Fattah, W., Reinhardt-Tews, A., Helm, M., Stark, M.J.R., Breunig, K.D., Schaffrath, R., Glatt, S.(2019) Nucleic Acids Res 47: 4814-4830
- PubMed: 30916349
- DOI: https://doi.org/10.1093/nar/gkz190
- Primary Citation of Related Structures:
6QP0 - PubMed Abstract:
Posttranscriptional RNA modifications occur in all domains of life. Modifications of anticodon bases are of particular importance for ribosomal decoding and proteome homeostasis. The Elongator complex modifies uridines in the wobble position and is highly conserved in eukaryotes. Despite recent insights into Elongator's architecture, the structure and function of its regulatory factor Kti12 have remained elusive. Here, we present the crystal structure of Kti12's nucleotide hydrolase domain trapped in a transition state of ATP hydrolysis. The structure reveals striking similarities to an O-phosphoseryl-tRNA kinase involved in the selenocysteine pathway. Both proteins employ similar mechanisms of tRNA binding and show tRNASec-dependent ATPase activity. In addition, we demonstrate that Kti12 binds directly to Elongator and that ATP hydrolysis is crucial for Elongator to maintain proper tRNA anticodon modification levels in vivo. In summary, our data reveal a hitherto uncharacterized link between two translational control pathways that regulate selenocysteine incorporation and affect ribosomal tRNA selection via specific tRNA modifications.
Organizational Affiliation:
Max Planck Research Group at the Malopolska Centre of Biotechnology, Jagiellonian University, Krakow, Poland.