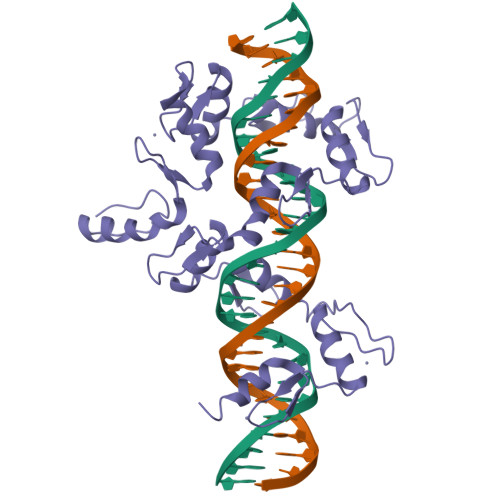
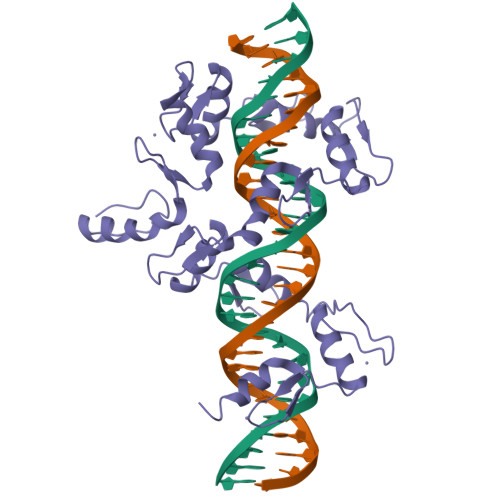
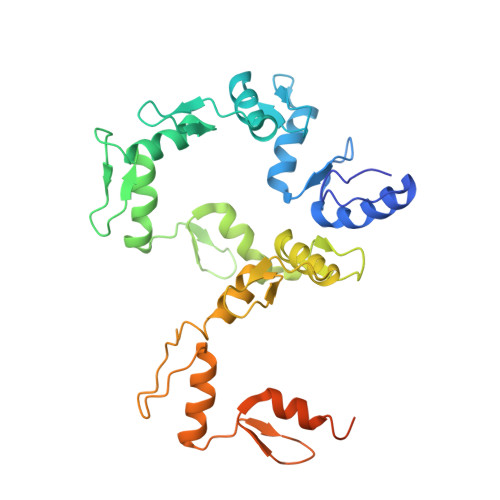
DNA Conformation Induces Adaptable Binding by Tandem Zinc Finger Proteins.
Patel, A., Yang, P., Tinkham, M., Pradhan, M., Sun, M.A., Wang, Y., Hoang, D., Wolf, G., Horton, J.R., Zhang, X., Macfarlan, T., Cheng, X.(2018) Cell 173: 221-233.e12
- PubMed: 29551271
- DOI: https://doi.org/10.1016/j.cell.2018.02.058
- Primary Citation of Related Structures:
5V3J, 5V3M, 5WJQ - PubMed Abstract:
Tandem zinc finger (ZF) proteins are the largest and most rapidly diverging family of DNA-binding transcription regulators in mammals. ZFP568 represses a transcript of placental-specific insulin like growth factor 2 (Igf2-P0) in mice. ZFP568 binds a 24-base pair sequence-specific element upstream of Igf2-P0 via the eleven-ZF array. Both DNA and protein conformations deviate from the conventional one finger-three bases recognition, with individual ZFs contacting 2, 3, or 4 bases and recognizing thymine on the opposite strand. These interactions arise from a shortened minor groove caused by an AT-rich stretch, suggesting adaptability of ZF arrays to sequence variations. Despite conservation in mammals, mutations at Igf2 and ZFP568 reduce their binding affinity in chimpanzee and humans. Our studies provide important insights into the evolutionary and structural dynamics of ZF-DNA interactions that play a key role in mammalian development and evolution.
Organizational Affiliation:
Department of Biochemistry, Emory University School of Medicine, 1510 Clifton Road, Atlanta, GA 30322, USA.