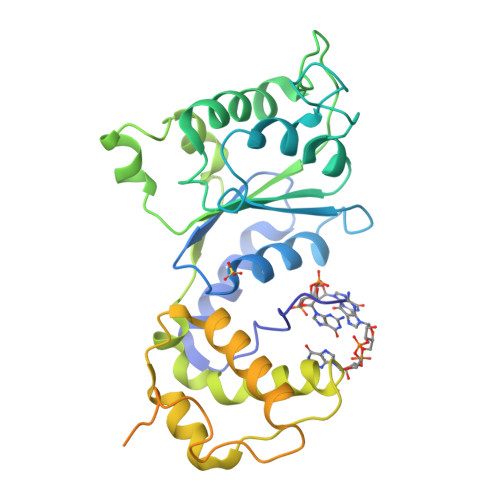
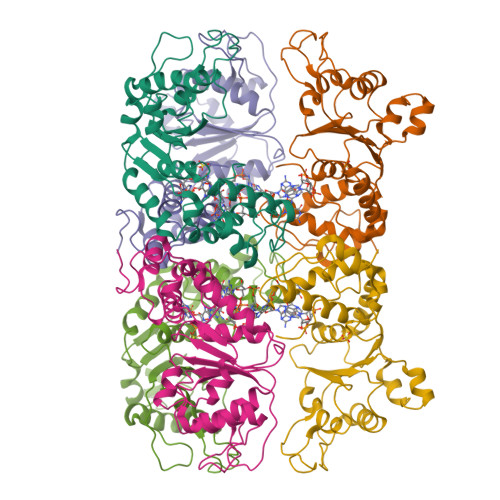
Mechanistic insights into c-di-GMP-dependent control of the biofilm regulator FleQ from Pseudomonas aeruginosa.
Matsuyama, B.Y., Krasteva, P.V., Baraquet, C., Harwood, C.S., Sondermann, H., Navarro, M.V.(2016) Proc Natl Acad Sci U S A 113: E209-E218
- PubMed: 26712005
- DOI: https://doi.org/10.1073/pnas.1523148113
- Primary Citation of Related Structures:
5EXP, 5EXS, 5EXT, 5EXX - PubMed Abstract:
Bacterial biofilm formation during chronic infections confers increased fitness, antibiotic tolerance, and cytotoxicity. In many pathogens, the transition from a planktonic lifestyle to collaborative, sessile biofilms represents a regulated process orchestrated by the intracellular second-messenger c-di-GMP. A main effector for c-di-GMP signaling in the opportunistic pathogen Pseudomonas aeruginosa is the transcription regulator FleQ. FleQ is a bacterial enhancer-binding protein (bEBP) with a central AAA+ ATPase σ(54)-interaction domain, flanked by a C-terminal helix-turn-helix DNA-binding motif and a divergent N-terminal receiver domain. Together with a second ATPase, FleN, FleQ regulates the expression of flagellar and exopolysaccharide biosynthesis genes in response to cellular c-di-GMP. Here we report structural and functional data that reveal an unexpected mode of c-di-GMP recognition that is associated with major conformational rearrangements in FleQ. Crystal structures of FleQ's AAA+ ATPase domain in its apo-state or bound to ADP or ATP-γ-S show conformations reminiscent of the activated ring-shaped assemblies of other bEBPs. As revealed by the structure of c-di-GMP-complexed FleQ, the second messenger interacts with the AAA+ ATPase domain at a site distinct from the ATP binding pocket. c-di-GMP interaction leads to active site obstruction, hexameric ring destabilization, and discrete quaternary structure transitions. Solution and cell-based studies confirm coupling of the ATPase active site and c-di-GMP binding, as well as the functional significance of crystallographic interprotomer interfaces. Taken together, our data offer unprecedented insight into conserved regulatory mechanisms of gene expression under direct c-di-GMP control via FleQ and FleQ-like bEBPs.
Organizational Affiliation:
Departamento de Física e Ciência Interdisciplinar, Instituto de Física de São Carlos, Universidade de São Paulo, São Carlos 13563-120, SP, Brazil;