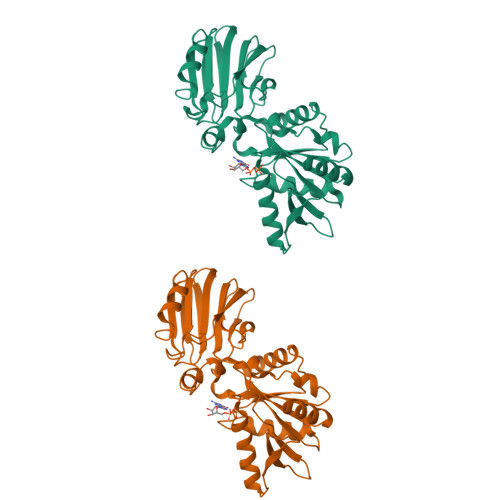
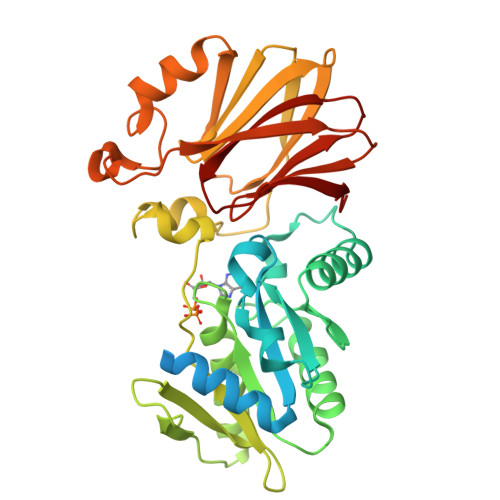
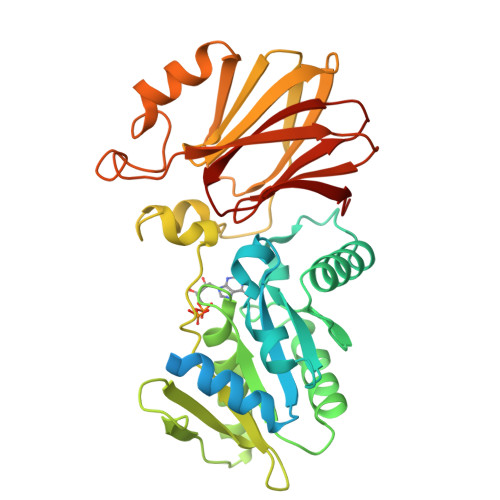
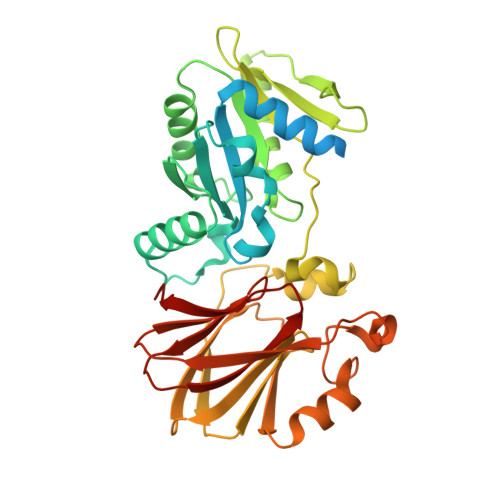
The structure of YqeH. An AtNOS1/AtNOA1 ortholog that couples GTP hydrolysis to molecular recognition.
Sudhamsu, J., Lee, G.I., Klessig, D.F., Crane, B.R.(2008) J Biological Chem 283: 32968-32976
- PubMed: 18801747
- DOI: https://doi.org/10.1074/jbc.M804837200
- Primary Citation of Related Structures:
3EC1 - PubMed Abstract:
AtNOS1/AtNOA1 was identified as a nitric oxide-generating enzyme in plants, but that function has recently been questioned. To resolve issues surrounding AtNOA1 activity, we report the biochemical properties and a 2.36 A resolution crystal structure of a bacterial AtNOA1 ortholog (YqeH). Geobacillus YqeH fused to a putative AtNOA1 leader peptide complements growth and morphological defects of Atnoa1 mutant plants. YqeH does not synthesize nitric oxide from L-arginine but rather hydrolyzes GTP. The YqeH structure reveals a circularly permuted GTPase domain and an unusual C-terminal beta-domain. A small N-terminal domain, disordered in the structure, binds zinc. Structural homology among the C-terminal domain, the RNA-binding regulator TRAP, and the hypoxia factor pVHL define a recognition module for peptides and nucleic acids. TRAP residues important for RNA binding are conserved by the YqeH C-terminal domain, whose positioning is coupled to GTP hydrolysis. YqeH and AtNOA1 probably act as G-proteins that regulate nucleic acid recognition and not as nitric-oxide synthases.
Organizational Affiliation:
Department of Chemistry and Chemical Biology, Cornell University, Ithaca, New York 14853, USA.