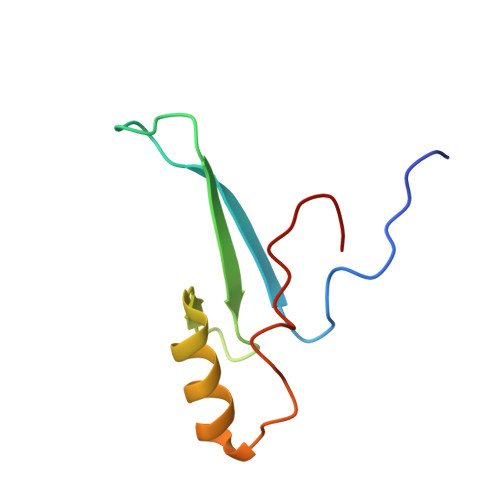
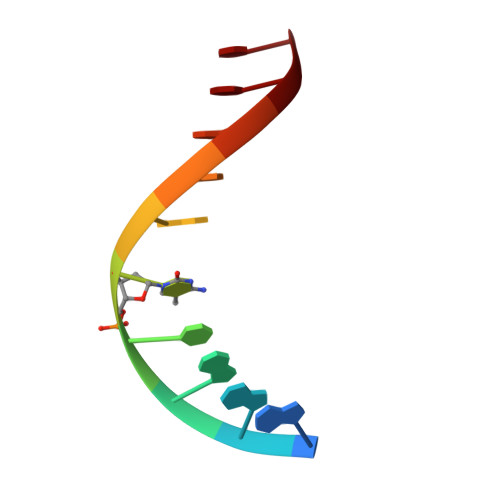
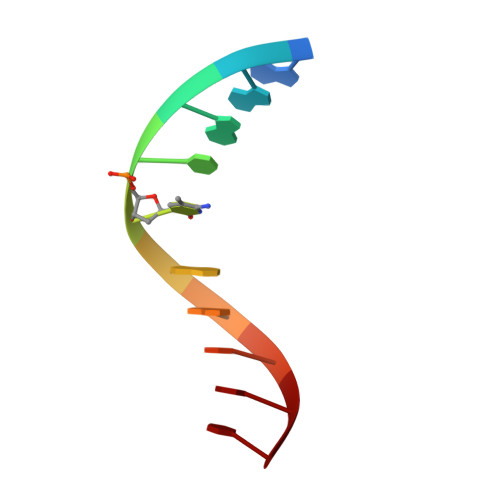
Solution structure and intramolecular exchange of methyl-cytosine binding domain protein 4 (MBD4) on DNA suggests a mechanism to scan for mCpG/TpG mismatches.
Walavalkar, N.M., Cramer, J.M., Buchwald, W.A., Scarsdale, J.N., Williams, D.C.(2015) Nucleic Acids Res 42: 11218-11232
- PubMed: 25183517
- DOI: https://doi.org/10.1093/nar/gku782
- Primary Citation of Related Structures:
2MOE - PubMed Abstract:
Unlike other members of the methyl-cytosine binding domain (MBD) family, MBD4 serves as a potent DNA glycosylase in DNA mismatch repair specifically targeting mCpG/TpG mismatches arising from spontaneous deamination of methyl-cytosine. The protein contains an N-terminal MBD (MBD4MBD) and a C-terminal glycosylase domain (MBD4GD) separated by a long linker. This arrangement suggests that the MBD4MBD either directly augments enzymatic catalysis by the MBD4GD or targets the protein to regions enriched for mCpG/TpG mismatches. Here we present structural and dynamic studies of MBD4MBD bound to dsDNA. We show that MBD4MBD binds with a modest preference for mCpG as compared to mismatch, unmethylated and hydroxymethylated DNA. We find that while MBD4MBD exhibits slow exchange between molecules of DNA (intermolecular exchange), the domain exhibits fast exchange between two sites in the same molecule of dsDNA (intramolecular exchange). Introducing a single-strand defect between binding sites does not greatly reduce the intramolecular exchange rate, consistent with a local hopping mechanism for moving along the DNA. These results support a model in which the MBD4MBD4 targets the intact protein to (m)CpG islands and promotes scanning by rapidly exchanging between successive mCpG sites which facilitates repair of nearby mCpG/TpG mismatches by the glycosylase domain.
Organizational Affiliation:
Department of Pathology and Laboratory Medicine, University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA.