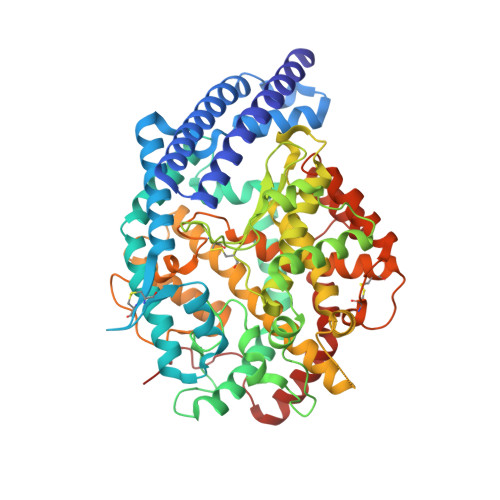
Molecular Recognition and Regulation of Human Angiotensin-I Converting Enzyme (Ace) Activity by Natural Inhibitory Peptides.
Masuyer, G., Schwager, S.L.U., Sturrock, E.D., Isaac, R.E., Acharya, K.R.(2012) Sci Rep 2: 717
- PubMed: 23056909
- DOI: https://doi.org/10.1038/srep00717
- Primary Citation of Related Structures:
4APH, 4APJ - PubMed Abstract:
Angiotensin-I converting enzyme (ACE), a two-domain dipeptidylcarboxypeptidase, is a key regulator of blood pressure as a result of its critical role in the renin-angiotensin-aldosterone and kallikrein-kinin systems. Hence it is an important drug target in the treatment of cardiovascular diseases. ACE is primarily known for its ability to cleave angiotensin I (Ang I) to the vasoactive octapeptide angiotensin II (Ang II), but is also able to cleave a number of other substrates including the vasodilator bradykinin and N-acetyl-Ser-Asp-Lys-Pro (Ac-SDKP), a physiological modulator of hematopoiesis. For the first time we provide a detailed biochemical and structural basis for the domain selectivity of the natural peptide inhibitors of ACE, bradykinin potentiating peptide b and Ang II. Moreover, Ang II showed selective competitive inhibition of the carboxy-terminal domain of human somatic ACE providing evidence for a regulatory role in the human renin-angiotensin system (RAS).
Organizational Affiliation:
Department of Biology and Biochemistry, University of Bath, Claverton Down, Bath, UK.