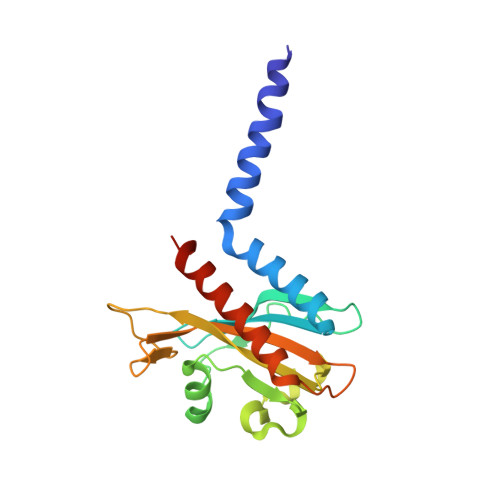
Crystal structure of a far-red-sensing cyanobacteriochrome reveals an atypical bilin conformation and spectral tuning mechanism.
Bandara, S., Rockwell, N.C., Zeng, X., Ren, Z., Wang, C., Shin, H., Martin, S.S., Moreno, M.V., Lagarias, J.C., Yang, X.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 33727422 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2025094118
- Primary Citation Related Structures:
6UV8, 6UVB - PubMed Abstract:
Cyanobacteriochromes (CBCRs) are small, linear tetrapyrrole (bilin)-binding photoreceptors in the phytochrome superfamily that regulate diverse light-mediated adaptive processes in cyanobacteria. More spectrally diverse than canonical red/far-red-sensing phytochromes, CBCRs were thought to be restricted to sensing visible and near UV light until recently when several subfamilies with far-red-sensing representatives (frCBCRs) were discovered. Two of these frCBCRs subfamilies have been shown to incorporate bilin precursors with larger pi-conjugated chromophores, while the third frCBCR subfamily uses the same phycocyanobilin precursor found in the bulk of the known CBCRs. To elucidate the molecular basis of far-red light perception by this third frCBCR subfamily, we determined the crystal structure of the far-red-absorbing dark state of one such frCBCR Anacy_2551g3 from Anabaena cylindrica PCC 7122 which exhibits a reversible far-red/orange photocycle. Determined by room temperature serial crystallography and cryocrystallography, the refined 2.7-Å structure reveals an unusual all-Z,syn configuration of the phycocyanobilin (PCB) chromophore that is considerably less extended than those of previously characterized red-light sensors in the phytochrome superfamily. Based on structural and spectroscopic comparisons with other bilin-binding proteins together with site-directed mutagenesis data, our studies reveal protein-chromophore interactions that are critical for the atypical bathochromic shift. Based on these analyses, we propose that far-red absorption in Anacy_2551g3 is the result of the additive effect of two distinct red-shift mechanisms involving cationic bilin lactim tautomers stabilized by a constrained all-Z,syn c onformation and specific interactions with a highly conserved anionic residue.
- Department of Chemistry, University of Illinois, Chicago, IL 60607.
Organizational Affiliation: