Searching for Cyclin-Dependent Kinase Inhibitors Using a New Variant of the Cope Elimination.
Griffin, R.J., Henderson, A., Curtin, N.J., Echalier, A., Endicott, J.A., Hardcastle, I.R., Newell, D.R., Noble, M.E., Wang, L.Z., Golding, B.T.(2006) J Am Chem Soc 128: 6012-6013
- PubMed: 16669651
- DOI: https://doi.org/10.1021/ja060595j
- Primary Citation of Related Structures:
2G9X - PubMed Abstract:
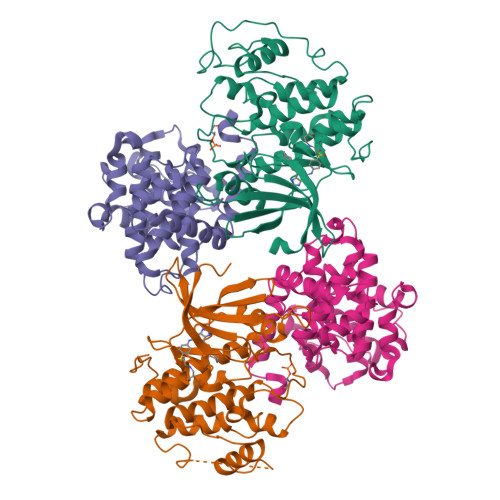
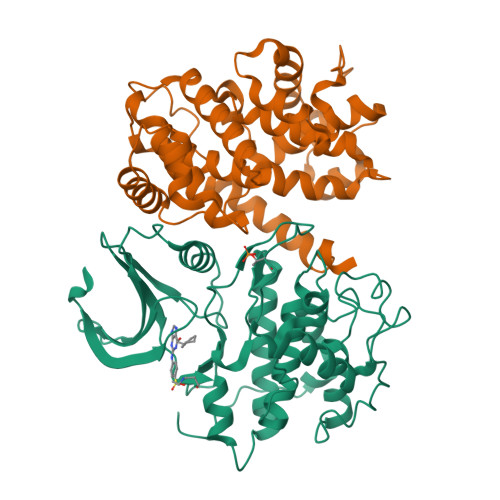
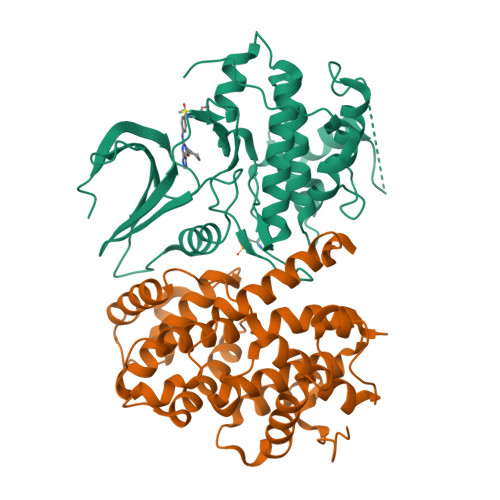
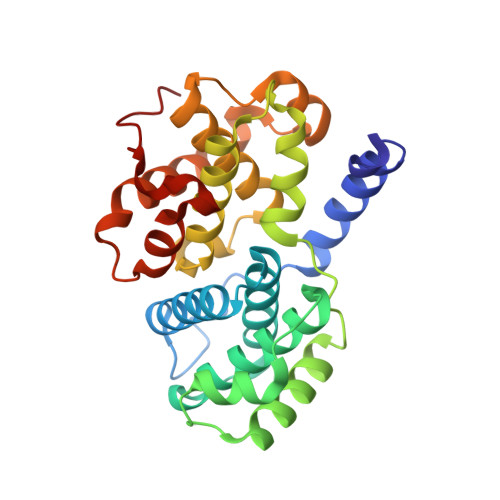
beta-Piperidinoethylsulfides are oxidized by m-chloroperbenzoic acid to intermediates containing both N-oxide and sulfone functions. These undergo a Cope-type elimination to a vinylsulfone that can be captured by amines to afford beta-aminoethylsulfones. When a beta-aminoethylsulfone group is linked to the 4-position of a phenyl group attached at N-2 of O6-cyclohexylmethylguanine, the resulting derivatives are inhibitors of the cyclin-dependent kinase CDK2. One of the most potent inhibitors (IC50 = 45 nM) contained a N-3-hydroxypropyl group on the aminoethylsulfonyl substituent. The crystal structure of this inhibitor bound to CDK2/cyclin A was determined and shows an unusual network of hydrogen bonds. The synthetic methodology developed can be utilized in multiple-parallel format and has numerous potential applications in medicinal chemistry.
Organizational Affiliation:
Northern Institute for Cancer Research, School of Natural Sciences-Chemistry, Bedson Building, University of Newcastle, Newcastle upon Tyne, UK.