JNJ-79032421, a Novel Membrane-restricted Mesothelin-targeting T-cell-engaging Bispecific Antibody for the Treatment of Mesothelin-positive Cancers.
Smans, K., Nesspor, T., De Breucker, S., van Heerde, E., Valckx, A., Van Slycken, N., Sproesser, K., Mattson Cypert, B., Packman, K., Chu, G., Fisher, J., Mazzanti, N., Petley, T., Chan, W., Del Rosario, B., Liu, T., Ember, S., Shaffer, P.L., Clawson, J.S., Bauml, J.M., Moores, S., Laquerre, S., Gavine, P.R.(2026) Mol Cancer Ther
- PubMed: 41773493
- DOI: https://doi.org/10.1158/1535-7163.MCT-25-0665
- Primary Citation Related Structures:
9Z3B - PubMed Abstract:
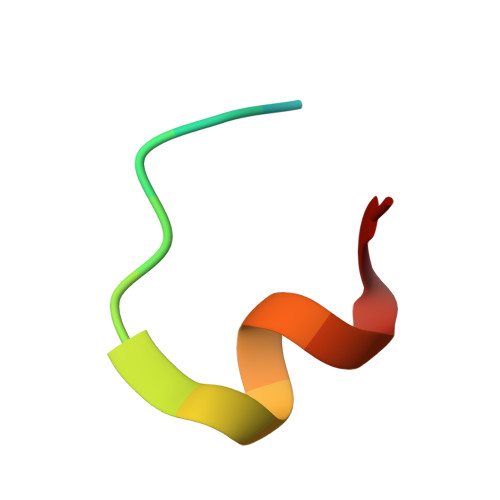
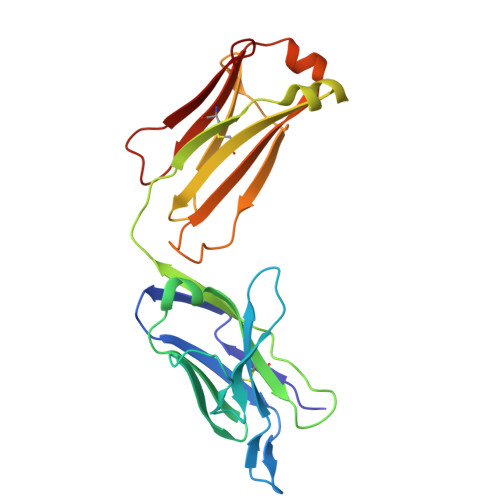
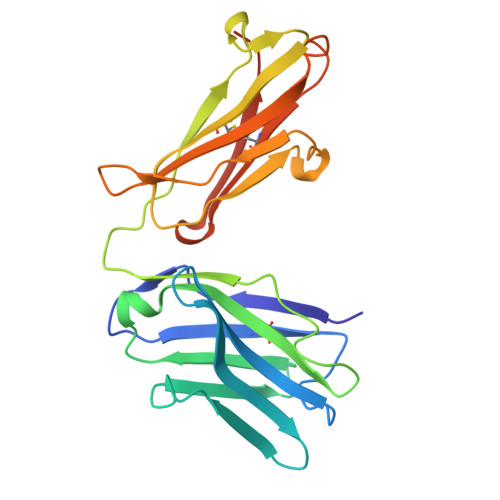
Mesothelin (MSLN) is a cell-membrane-anchored glycoprotein overexpressed in several cancers including pancreatic cancer, ovarian cancer, and mesothelioma. MSLN is protease-cleaved at the membrane-proximal region, releasing shed MSLN (sMSLN) into the tumor microenvironment, leaving behind a membrane-bound stub. sMSLN in the tumor microenvironment is hypothesized to create a 'sink' that could limit tumor binding of MSLN-targeted anticancer therapies. We describe JNJ-79032421, a bispecific antibody targeting cluster of differentiation (CD)3 on T cells and the membrane-restricted, non-shed region of full-length MSLN on cancer cells. Crystal structure analysis confirmed JNJ-79032421 binding to the membrane-restricted, non-shed C-terminal region of MSLN. Using Western blot, enzyme-linked immunosorbent assay, fluorescence-activated cell sorting, and immunohistochemistry, elevated levels of sMSLN were detected in blood, serosal fluids, and tumor tissue of patients and tended to increase with MSLN tumor expression. Binding and cytotoxicity of JNJ-79032421 against cancer cell lines was unaffected by the presence of sMSLN. In cancer cell lines and cell-line-derived and patient-derived xenograft mouse models, increasing levels of MSLN expression led to increased JNJ-79032421 potency. In contrast, MSLNxCD3 bispecific antibodies whose binding is non membrane restricted demonstrated a loss of cytotoxicity in the presence of sMSLN. This 'sink' effect was most pronounced for high-affinity non-membrane-restricted bispecific antibodies. Increases of CD45+ and CD8+ T-cell infiltrates, as well as CD8+ T-cell activation, observed in xenograft tumor tissues were time and JNJ-79032421 dose dependent. Overall, these data suggest that JNJ-79032421 may offer meaningful clinical activity by avoiding binding to sMSLN.
- Johnson & Johnson (United States) Beerse Belgium.
Organizational Affiliation: