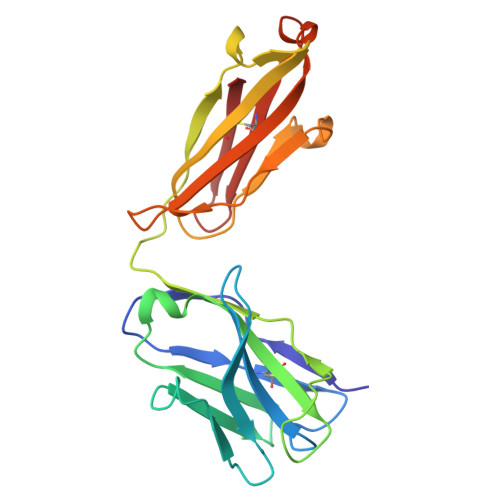
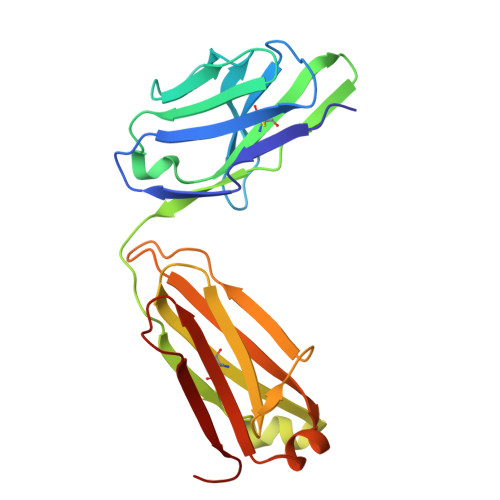
Targeting peptide-MHC complexes with designed T cell receptors and antibodies.
Motmaen, A., Jude, K.M., Wang, N., Minervina, A., Feldman, D., Lichtenstein, M.A., Ebenezer, A., Correnti, C., Thomas, P.G., Garcia, K.C., Baker, D., Bradley, P.(2025) bioRxiv
- PubMed: 41332722 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2025.11.19.689381
- Primary Citation Related Structures:
9YTD, 9YTF - PubMed Abstract:
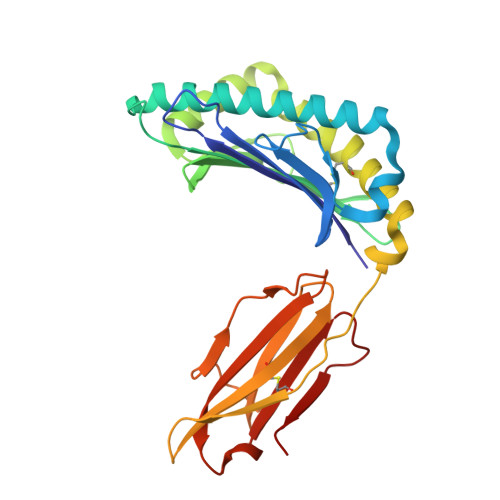
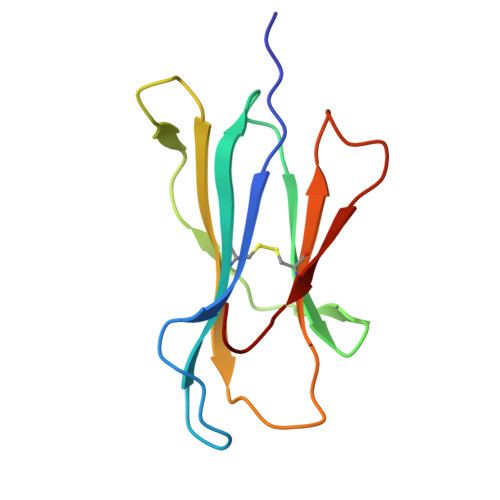
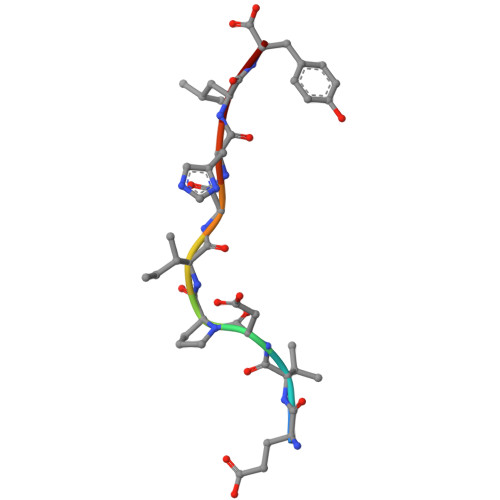
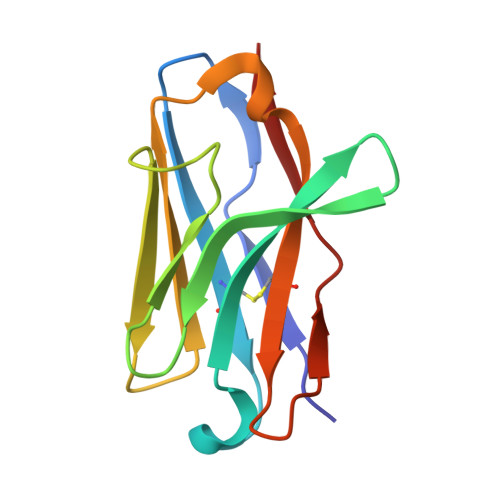
Class I major histocompatibility complexes (MHCs), expressed on the surface of all nucleated cells, present peptides derived from intracellular proteins for surveillance by T cells. The precise recognition of foreign or mutated peptide-MHC (pMHC) complexes by T cell receptors (TCRs) is central to immune defense against pathogens and tumors. Although patient-derived TCRs specific for cancer-associated antigens have been used to engineer tumor-targeting therapies, their reactivity toward self- or near-self antigens may be constrained by negative selection in the thymus. Here, we introduce a structure-based deep learning framework, ADAPT (Antigen-receptor Design Against Peptide-MHC Targets), for the design of TCRs and antibodies that bind to pMHC targets of interest. We evaluate the ADAPT pipeline by designing and characterizing TCRs and antibodies against a diverse panel of pMHCs. Cryogenic electron microscopy structures of two designed antibodies bound to their respective pMHC targets demonstrate atomic-level accuracy at the recognition interface, supporting the robustness of our structure-based approach. Computationally designed TCRs and antibodies targeting pMHC complexes could enable a broad range of therapeutic applications, from cancer immunotherapy to autoimmune disease treatment, and insights gained from TCR-pMHC design should advance predictive understanding of TCR specificity with implications for basic immunology and clinical diagnostics.
- Institute for Protein Design, University of Washington, Seattle, WA, USA.
Organizational Affiliation: