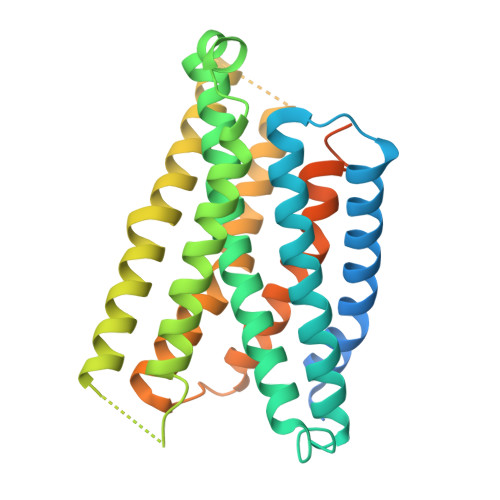
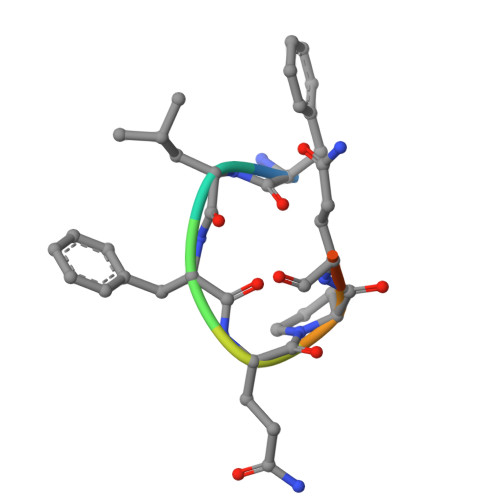
Structural insight into ligand binding and activation of the orphan GPCR Mas1.
Zhang, Y., Wang, Q., Liu, H., Shan, H., Gu, Y., Yang, J., Gao, Y., Wu, K., Yang, D., Xu, H.E.(2026) EMBO J 45: 3500-3513
- PubMed: 41912627 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s44318-026-00764-6
- Primary Citation Related Structures:
9X3Y, 9X3Z, 9X40, 9X41 - PubMed Abstract:
The Mas1 receptor, an orphan class A G-protein-coupled receptor (GPCR), plays pivotal roles in cardiovascular and anti-inflammatory regulation. Despite its therapeutic relevance, the structural mechanisms underlying Mas1 ligand binding and activation remain poorly understood. Here, we report cryo-EM structures of Mas1 bound to two chemically distinct agonists-neuropeptide FF (NPFF) and synthetic small-molecule AR234958-captured in complex with inhibitory G proteins. These structures reveal a conserved orthosteric binding pocket accommodating both ligands through shared hydrophobic interactions. Unlike many other class A GPCRs that rely on direct W 6.48 toggle switch engagement, Mas1 adopts a non-canonical activation strategy driven by a ligand-induced hydrophobic compression plane involving residues Y248 6.55 , L87 2.60 , I84 2.57 , and L266 7.39 at the bottom of the ligand binding pocket. This mechanism transmits mechanical tension to promote TM6 displacement and G protein coupling. Functional mutagenesis validates this model, identifying two transmembrane helix 6 (TM6) residues, M244 6.51 and F237 6.44 , as critical molecular switches. Comparative analyses of Mas1-related receptors, MRGPRX1-X4, reveal conserved features and mechanistic divergence within this subfamily. These findings provide a structural framework for understanding Mas1 pharmacology and rational design of selective therapeutics.
- State Key Laboratory of Drug Research, Center for Structure and Function of Drug Targets, Shanghai Institute of Materia Medica, Chinese Academy of Sciences, Shanghai, China. yumu_zhang@hms.harvard.edu.
Organizational Affiliation: