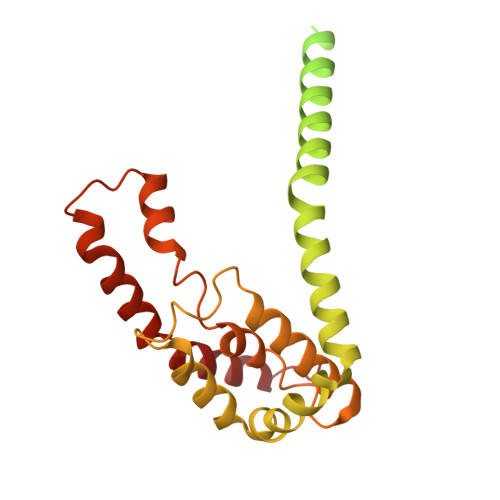
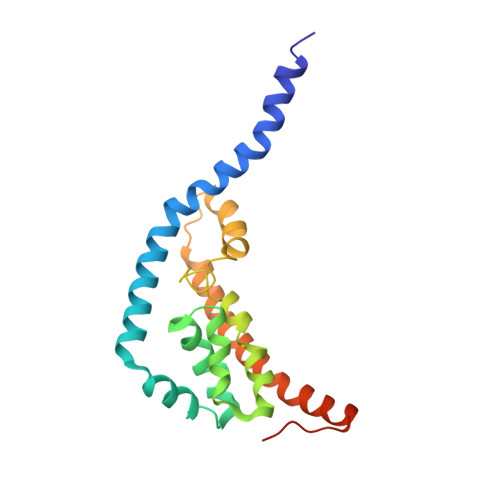
A trimeric architecture reveals the glucitol PTS transporter as a distinct superfamily.
Deng, T., Liu, X., Zeng, J., Ge, X., Wang, J.(2026) Commun Biol 9
- PubMed: 41807737 Search on PubMed
- DOI: https://doi.org/10.1038/s42003-026-09835-0
- Primary Citation Related Structures:
9X2M - PubMed Abstract:
The bacterial phosphoenolpyruvate:sugar phosphotransferase system (PTS) catalyzes the transport and phosphorylation of carbohydrates. The glucitol (Gut) PTS transporter from Escherichia coli has often been discussed in relation to the Glucose-Fructose-Lactose (GFL) superfamily, although other work has suggested that it may instead form a separate PTS superfamily. This uncertainty is linked to its unusual genetic organization, in which the transmembrane IIC domain is divided into two polypeptides (IIC1/GutE and IIC2/GutA). Here, we present the cryo-electron microscopy (cryo-EM) structure of the complete Gut transporter, which resolves this discrepancy by revealing a homotrimeric architecture for its transmembrane domain-a fold unprecedented among sugar-transporting PTS permeases. This structural evidence supports the view that the Gut family represents a distinct PTS superfamily. Within the trimer, the protomers are captured in inward-facing and inward-occluded conformations, providing a structural basis for an alternating-access transport mechanism. Furthermore, the structure suggests a unique in-trans phosphotransfer pathway between the IIB and IIC domains of adjacent subunits and identifies the substrate-binding pocket at the GutA/GutE interface. Our work redefines the structural landscape of PTS transporters and provides a mechanistic framework for sugar transport by this unique trimeric porter.
- State Key Laboratory of Membrane Biology, Beijing Frontier Research Center for Biological Structure, School of Life Sciences, Tsinghua University, Beijing, China.
Organizational Affiliation: