Structural insights into sigma 28-dependent transcription initiation and its regulation by anti-sigma factor in Pseudomonas aeruginosa
Sheenu, N., Kumar, V., Sahoo, P.K., Kandiah, E., Jain, D.(2026) Nucleic Acids Res 54
- PubMed: 41521663 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaf1414
- Primary Citation Related Structures:
9WSF, 9WSM - PubMed Abstract:
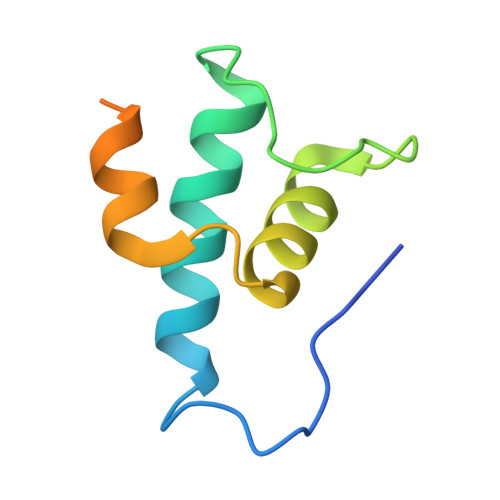
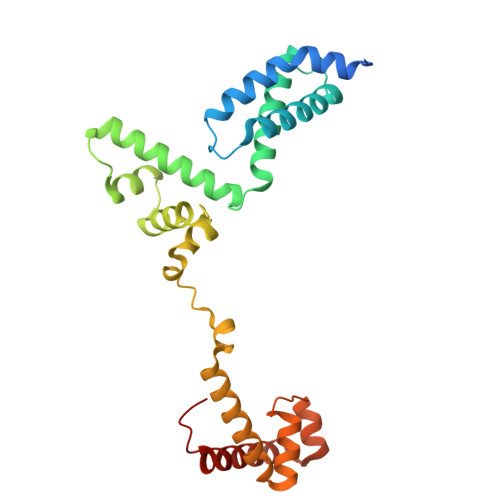
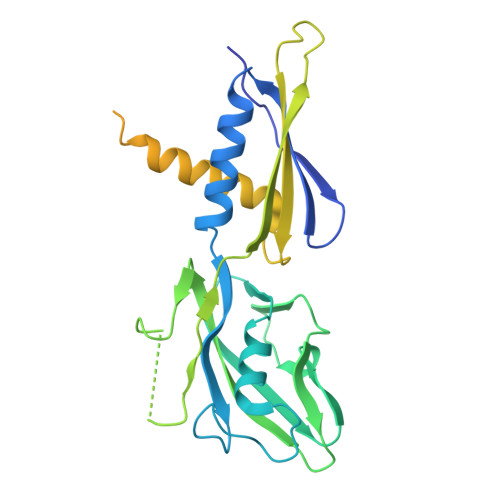
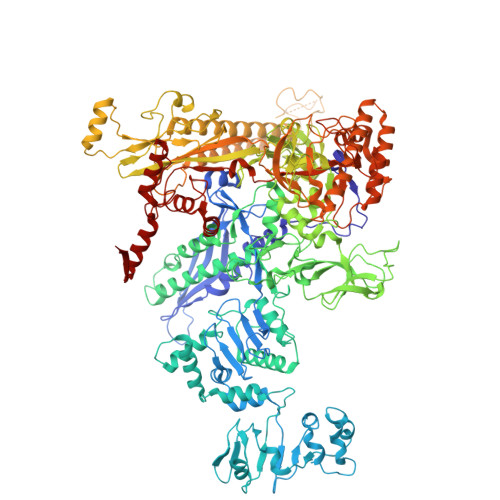
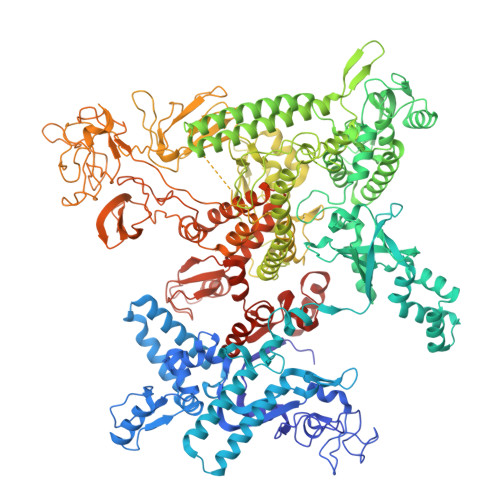
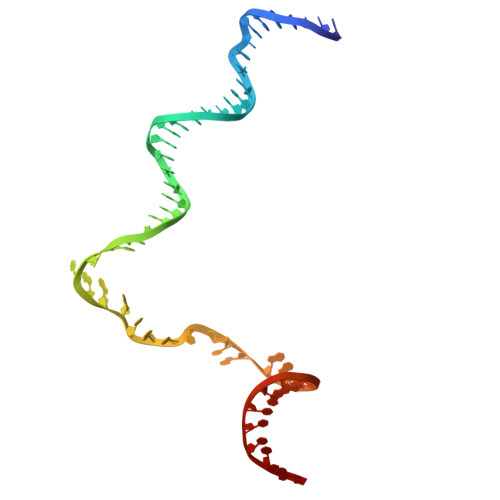
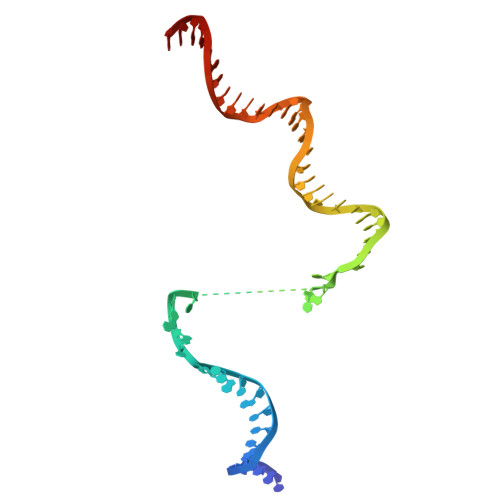
Late flagellar genes in Pseudomonas aeruginosa are transcribed by the group 3 sigma factor, FliA (σ28). σ28 drives the expression of flagellin, which assembles into the flagellar filament in this monoflagellated bacterium. This function is suppressed by the anti-sigma factor FlgM. Here, we present the 1.95Å resolution crystal structure of σ28-FlgM complex, along with a 3.4Å structure of σ28RNAP open promoter complex determined using single particle cryo-electron microscopy from P. aeruginosa. The σ28 adopts a compact conformation upon binding to the anti-sigma factor FlgM, which contacts all three domains of the sigma factor. This conformation is neither conducive to interactions with RNA polymerase nor the promoter DNA. The cryo-EM structure reveals base-specific interactions of σ28 domain 4 (σ4) with -35 element, flipping of -11 base of the template strand, novel interactions of template strand with domain 2 (σ2) and 3 (σ3), and partial insertion of sigma finger into the active site cleft, offering unique features of group 3 sigma interactions with promoter DNA. Perturbation of key residues affects transcription in vitro and flagellar phenotypes as well as bacterial motility in vivo. Analysis of the structural data presented here reveals new insights into transcription regulation of late flagellar genes.
- Transcription Regulation Lab, Regional Centre for Biotechnology, NCR Biotech Science Cluster, 3rd Milestone, Faridabad-Gurgaon Expressway, Faridabad 121001, India.
Organizational Affiliation: