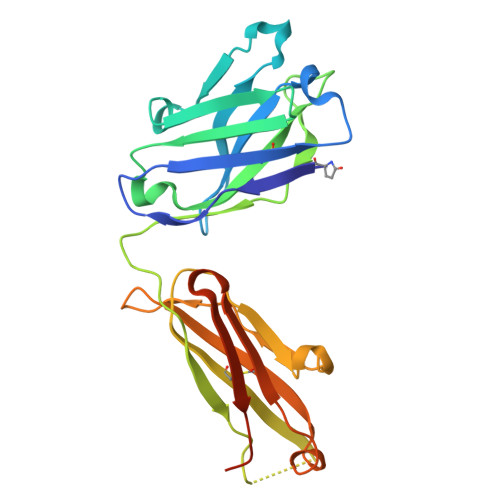
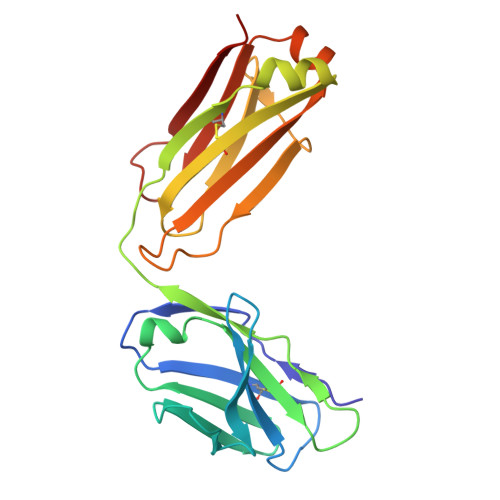
Structural basis of B-to-Z DNA transition mediated by an anti-Z-DNA antibody.
Lee, C.C., Hsu, S.F., Chang, Y.W., Chen, Y.W., Ho, M.R., Sugiyama, H., Wang, W.C., Wang, A.H.(2026) Nucleic Acids Res 54
- PubMed: 41761907 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkag160
- Primary Citation Related Structures:
9WRN, 9WS0, 9WU2 - PubMed Abstract:
Z-DNA is a left-handed double-helical form of DNA that plays roles in transcription, immune responses, viral infection, bacterial biofilm formation, and autoimmune diseases. Despite its importance, the instability of Z-DNA under physiological conditions has hindered detailed structural and functional investigations. Moreover, although antibodies are known to recognize nucleic acids, the mechanisms underlying their detection and stabilization of dynamic DNA under biological conditions remain unclear. This study provides the first atomic-level structural insights into antibody-mediated B-to-Z DNA transition. Accordingly, a Z-DNA-specific chimeric Fab fragment of Z22 (cZ22-Fab) was designed and characterized using multiple biophysical approaches. cZ22-Fab mediates a concentration-dependent B-to-Z conformational transition in CG-repeat DNA, establishing a stable 2:1 Fab/DNA stoichiometry. Crystal structure of cZ22-Fab/Z-DNA complexes revealed a left-handed DNA backbone-tracking recognition mode, in which cZ22-Fab recognizes Z-DNA conformation through phosphate-clamping and base interactions. Notably, a 5'-end C-hanging Z-DNA duplex structure formed by dC(GC)3 and stabilized by cZ22-Fab was observed. Structure-guided mutagenesis demonstrated that heavy chain residues R50 and Y106 are critical for Z-DNA binding, and analyses of additional Z-DNA-forming sequences further elucidated the binding characteristics. Overall, this work provides molecular insights into the mechanism of antibody-mediated Z-DNA formation and stabilization, highlighting its therapeutic relevance and implications for autoimmunity.
- The Ph.D. Program for Translational Medicine, College of Medical Science and Technology, Taipei Medical University, Taipei 110301, Taiwan.
Organizational Affiliation: