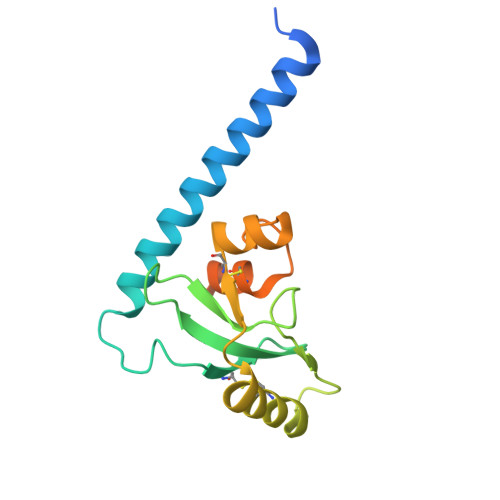
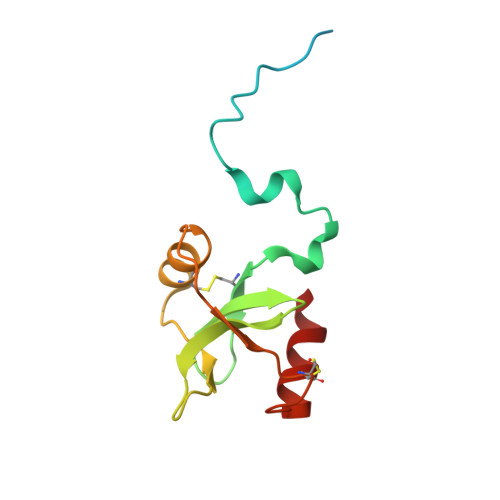
Structural basis of mpox virus A30/H2 subcomplex formation.
Jia, X., Lin, S., Yang, F., You, Y., Yang, R., Chen, Z., Guo, L., Yang, J., Wang, L., Yuan, X., Zhang, X., Xu, P., Tong, Q., He, B., Cao, Y., Li, J., Zhao, Q., Lu, G.(2026) Proc Natl Acad Sci U S A 123: e2524604123-e2524604123
- PubMed: 41642990 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2524604123
- Primary Citation Related Structures:
9L7W, 9WLP - PubMed Abstract:
The continuous spread of mpox disease caused by mpox virus (MPXV) has posed great threat to global public health. The postattachment membrane fusion process of MPXV is mediated by a multimeric protein machinery, termed as entry-fusion complex (EFC). Among EFC components, A30 and H2 are the earliest identified interaction pair and play important roles in virus entry. Here, we determine the crystal structure of MPXV A30/H2 subcomplex via the tandem-fusion strategy, and show that A30 undergoes large conformational rearrangements upon H2 binding. Structural analysis reveals extended intersubunit interface and highly conserved intermolecular interactions. In vitro binding data further clarify key residues and elements involved in the A30/H2 subcomplex formation. Finally, we show that the H2-A30 fusion protein, superior to A30 ectodomain alone or the ectodomain-mixture of H2+A30, can induce more potent neutralizing-antibody responses which could inhibit viral infection. These data provide valuable information for the understanding of poxvirus EFC assembly and the H2-A30-based immunogen design and optimization.
- Department of Emergency Medicine, State Key Laboratory of Biotherapy, West China Hospital, Sichuan University, Chengdu 610041, Sichuan, China.
Organizational Affiliation: