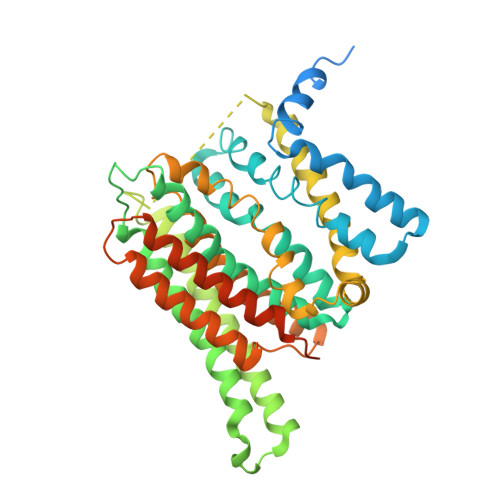
Structural basis of CAX1 autoinhibition by its amino-terminal domain in Arabidopsis thaliana.
Wang, K., Ma, C., Chen, G., Yang, Z., Gao, Y., Zhang, Z., Liu, X., Sun, L.(2025) Nat Plants 11: 2072-2083
- PubMed: 40897811 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41477-025-02104-8
- Primary Citation Related Structures:
9VSD - PubMed Abstract:
Calcium homeostasis is tightly regulated due to the essential roles of calcium ions (Ca 2+ ) in various cellular processes. CAX1 in Arabidopsis thaliana (AtCAX1) serves as a Ca 2+ /H + exchanger transporting excess cytosolic Ca 2+ into the vacuole, which is modulated by kinase phosphorylation in response to diverse signals. However, the regulatory mechanism remains unclear. Here we present the structures of wild-type AtCAX1 in an inactivated state and a phosphomimetic mutant in an activated state. In the wild-type structure, the amino-terminal region forms an α-helix that blocks the transport tunnel, thus inhibiting its transport activity. In contrast, in the phosphomimetic mutant structure, this blocking helix is released from the tunnel, leading to AtCAX1 activation. Conformational changes are also observed in the transmembrane domain. Together, these findings provide insights into the transport mechanism of the Ca 2+ /H + exchangers and set up a basis for future studies of the regulation of calcium homeostasis in plants.
- Department of Neurology, First Affiliated Hospital of USTC, MOE Key Laboratory for Membraneless Organelles and Cellular Dynamics, Hefei National Research Center for Physical Sciences at the Microscale, Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei, China.
Organizational Affiliation: