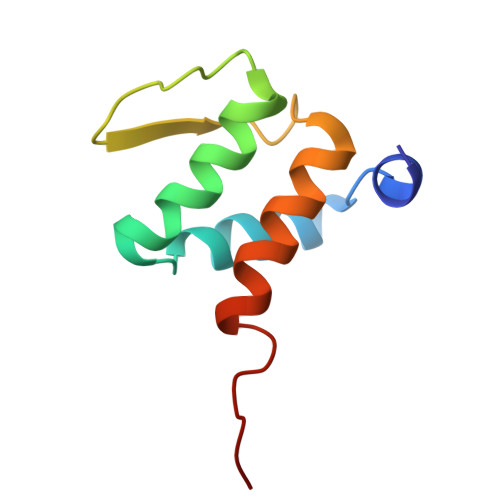
Recruitment of BRD4 to the ASXL1 genomic targets depends on the extra-terminal domain of BRD4.
Selvam, K., Lu, S., Messmer, C., Pang, Y., Biswas, S., Khalil, M., Zhang, P., Tulaiha, R., Zhou, M.M., Kitamura, T., Lauberth, S.M., Blanco, M.A., Yang, F.C., Affar, E.B., Zhao, Z., Zeng, L., Wang, L., Kutateladze, T.G.(2026) Nat Commun 17
- PubMed: 41702923
- DOI: https://doi.org/10.1038/s41467-026-69565-z
- Primary Citation Related Structures:
9VQ1 - PubMed Abstract:
ASXL1 is a well-established driver of a wide range of cancers. Here, we identify a high level of genetic correlation between ASXL1 and the major transcriptional activator BRD4 in cancer cells and characterize the molecular mechanism underlying this correlation. Structural and biochemical data show the formation of a tight complex between the extraterminal domain of BRD4 and the BRD4-binding motif of ASXL1. ChIP-seq analysis of mutated ASXL1 that cannot bind BRD4 demonstrates that the recruitment of BRD4 to the ASXL1 genomic targets depends on this interaction. We find that the cancer-related truncated variants comprising residues 1-645 and 1-591 of ASXL1 (ASXL1 1-645 and ASXL1 1-591 ) retain BRD4 binding function, with ASXL1 1-645 showing an enhanced ability to recruit BRD4 to promoters of ASXL1 target genes. In contrast to ASXL1 1-591 , ASXL1 1-645 simultaneously and independently interacts with BRD4 and the H3K4-specific methyltransferases MLL3/4, whereas the shorter variant is lacking the MLL3/4-binding site and interacts only with BRD4. Genomic data from six cancer types reveals a strong positive ASXL1-BRD4 relationship, with BRD4 occupying the ASXL1 promoter and thus pointing to a possible feed-forward mechanism. Our findings provide mechanistic details by which ASXL1 associates with BRD4 and shed light on the biological significance of this association.
- Department of Pharmacology, University of Colorado School of Medicine, Aurora, CO, USA.
Organizational Affiliation: