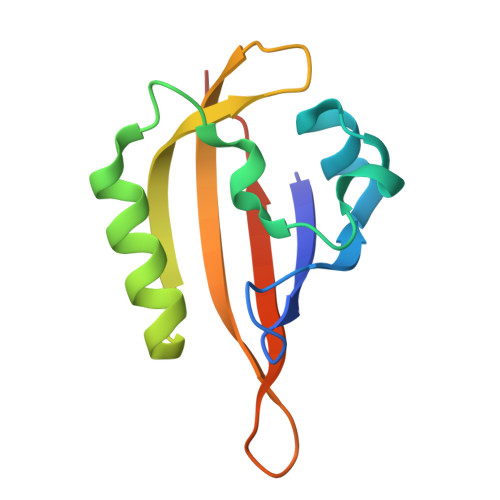
Structural characterization of Meiothermus ruber LOV domain.
Semenov, O., Nazarenko, V., Yudenko, A., Kovalev, K., Goncharov, I., Natarov, I., Mikhailov, A., Kuznetsova, E., Nikolaev, A., Yang, Y., Sluchanko, N.N., Borshchevskiy, V., Remeeva, A., Gushchin, I.(2025) J Struct Biol 218: 108268-108268
- PubMed: 41349808 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2025.108268
- Primary Citation Related Structures:
9VF7, 9VF8 - PubMed Abstract:
Light Oxygen Voltage (LOV) domains are important widespread receptors of blue light that also found applications in optogenetics and imaging. While LOV domains from mesophiles are relatively well characterized, their counterparts from thermophilic microorganisms remain understudied. Here, we express two constructs of a LOV domain belonging to a histidine kinase from Meiothermus ruber, MrLOV and MrLOVe, and show that they are photoactive, with recovery time values of 21 and 27 min, respectively, and thermostable. Crystal structures reveal that MrLOV, which lacks helices A'α and Jα, forms a parallel dimer, whereas MrLOVe is a tetramer organized as an antiparallel dimer of two parallel dimers interacting via helices Jα. One MrLOVe dimer is symmetric, and the other is asymmetric, with conformational differences mirroring activation-related changes in other LOV domains. Our data provide the structural basis for understanding and engineering of thermophilic LOVs and pave the way for development of thermostable and photostable LOV-derived optogenetic tools and flavin-based fluorescent proteins.
- Research Center for Molecular Mechanisms of Aging and Age-Related Diseases, Moscow Institute of Physics and Technology, 141700 Dolgoprudny, Russia.
Organizational Affiliation: