A user-friendly goniometer-compatible fixed-target platform for macromolecular crystallography at synchrotrons.
Ghosh, S., Banacore, A., Norder, P., Bjelcic, M., Kabbinale, A., Nileshwar, P., Wehlander, G., de Sanctis, D., Basu, S., Orlans, J., Vallejos, A., Chavas, L.M.G., Neutze, R., Branden, G.(2026) J Appl Crystallogr 59: 303-315
- PubMed: 41959872 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1600576725011513
- Primary Citation Related Structures:
9TBL, 9UYR, 9VDJ, 9VDX - PubMed Abstract:
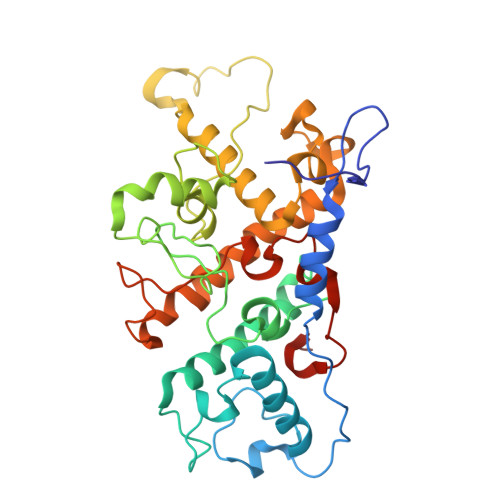
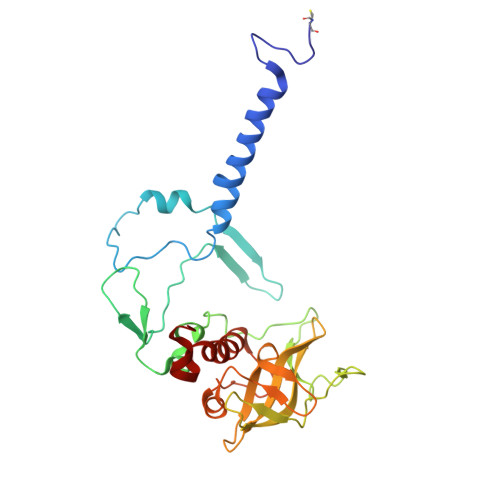
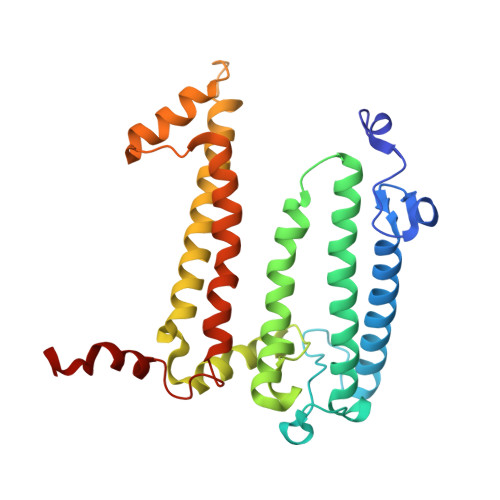
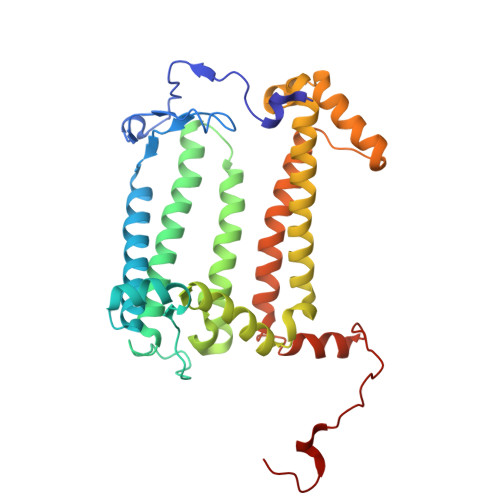
Fixed-target platforms provide convenient support for microcrystals during serial X-ray crystallography studies using synchrotron radiation. Here, we describe a simple user-friendly 3D-printed support where the crystals are sandwiched between two layers of thin X-ray-transparent membrane resulting in very low scattering background. The platform is compatible with magnetic mounting onto the standard goniometer of macromolecular crystallography beamlines. Our design utilizes a 96-well frame that facilitates hanging-drop experiments directly on the membrane using conventional crystallization plates, thereby eliminating multiple pipetting and crystal handling steps. Crystals can be enclosed in a sandwich and packed into 'cassettes', preventing the risk of the sample drying out during room-temperature transportation to synchrotron sources. The versatility of the platform is demonstrated by five structures solved using different crystallization and data-collection strategies. Lysozyme single-crystal rotational crystallography at room temperature is shown, as well as microcrystal serial data collection under cryogenic conditions. On-chip microcrystallization is illustrated by use of a photosynthetic reaction center as an example. Finally, serial crystallography data collection at room temperature from microcrystals of the membrane protein cytochrome c oxidase crystallized in lipidic cubic phase is presented.
- Department of Chemistry and Molecular Biology Gothenburg University Sweden.
Organizational Affiliation: