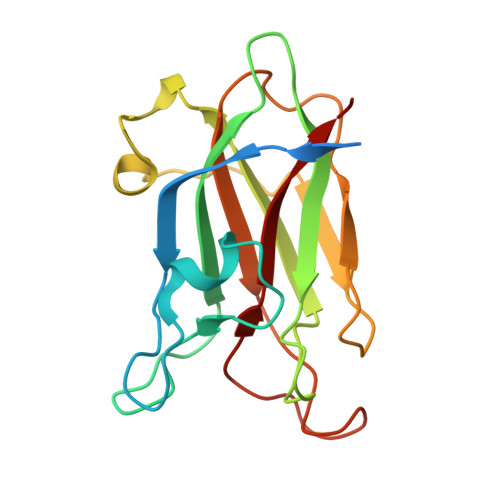
Structural insights into lacto-N-biose I recognition by a family 32 carbohydrate-binding module from Bifidobacterium bifidum.
Zhang, X., Sunagawa, N., Kashima, T., Igarashi, K., Miyanaga, A., Fushinobu, S.(2026) FEBS Lett 600: 504-514
- PubMed: 41204437 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.70217
- Primary Citation Related Structures:
9VAK, 9WFH - PubMed Abstract:
Bifidobacterium bifidum, a predominant colonizer of the infant gut, utilizes lacto-N-biose I (LNB), a prominent component of human milk oligosaccharides (HMOs), through a dedicated metabolic pathway. Among a diverse set of extracellular glycosidases involved in HMO degradation, lacto-N-biosidase (LnbB) plays a pivotal role by releasing LNB. We investigated the structure and function of the carbohydrate-binding module family 32 (CBM32) domain located at the C-terminus of the glycoside hydrolase family 20 catalytic domain in LnbB. Isothermal titration calorimetry showed that CBM32 binds LNB with a dissociation constant (K d ) of 98 μm. The crystal structure of the CBM32 complexed with LNB reveals the molecular basis for its specific recognition. Impact statement Bifidobacteria are beneficial gut microbes, and infant-associated strains establish symbiosis by degrading human milk oligosaccharides. This study uncovers the molecular mechanism by which Bifidobacterium bifidum captures lacto-N-biose I, a key disaccharide, functioning as a cross-feeder that promotes the growth of other bifidobacteria and supports the infant gut ecosystem.
- Department of Biotechnology, The University of Tokyo, Japan.
Organizational Affiliation: