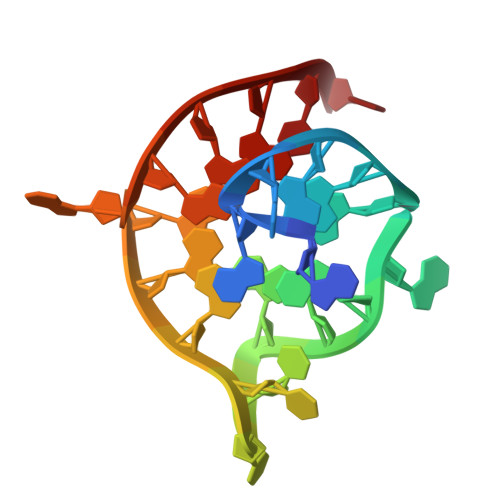
Dual targeting of topoisomerase I and DNA G-quadruplexes enhances senescence and chemosensitivity in colorectal cancer.
Li, Y., Ji, D., Jia, Y., Liang, Y., Wei, E., Gao, C., Li, Y., Zeng, C., Wang, L., Wang, C., Guo, Z., Zhang, Y., Zhou, M.M., Wu, D., Zeng, L.(2026) Commun Biol 9
- PubMed: 41803560 Search on PubMed
- DOI: https://doi.org/10.1038/s42003-026-09801-w
- Primary Citation Related Structures:
9VA0, 9VA3 - PubMed Abstract:
Colorectal cancer (CRC) remains a therapeutic challenge due to chemoresistance that limits conventional treatment efficacy. We developed ZBH-01, a camptothecin derivative engineered to target both topoisomerase I (TOP1) and DNA G-quadruplexes (G4s). Unlike irinotecan (CPT-11), which requires metabolic activation, ZBH-01 directly stabilizes TOP1-DNA covalent complexes and preferentially binds the hTERT promoter G4, a regulator of telomere maintenance and oncogenic transcription. Structural studies reveal that the crescent-shaped scaffold of ZBH-01 π-π stacks onto the external G-tetrad of the hTERT G4, displacing SP1/MYC transcription factors and suppressing hTERT expression. Functionally, ZBH-01 demonstrated improved efficacy in chemoresistant models, exhibiting 14-fold and 7-fold greater efficacy than CPT-11 and SN-38 respectively in cisplatin-resistant cells, and outperforming CPT-11 by 61-fold and SN-38 by 2.4-fold in 5-FU-resistant models. By concurrently disrupting DNA repair through TOP1-trapping and transcriptional adaptation via G4-stabilization, ZBH-01 induced DNA damage, telomere shortening, and cellular senescence. These findings establish TOP1/G4 dual-targeting as a potential therapeutic strategy to enhance CRC chemosensitivity, presenting a new framework for combining DNA damage induction with transcription modulation.
- Institute of Translational Medicine, The First Hospital of Jilin University, Changchun, China.
Organizational Affiliation: