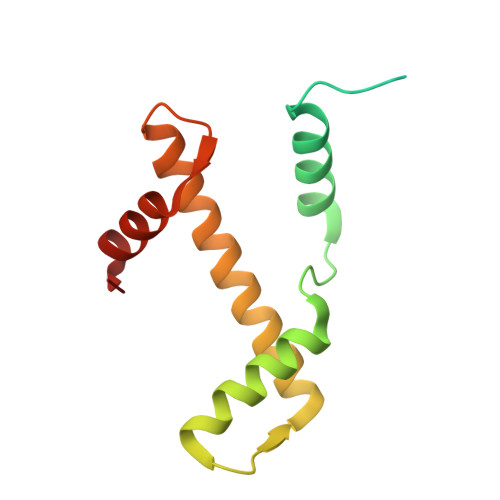
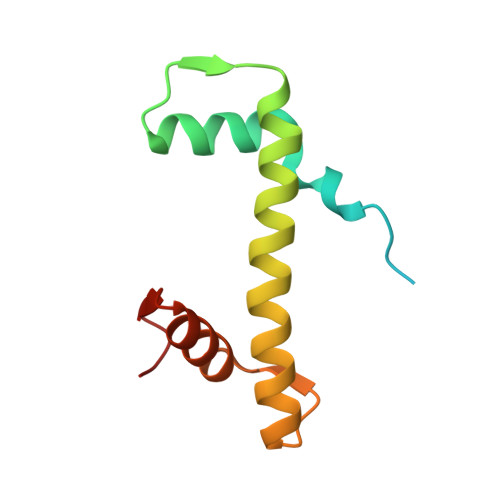
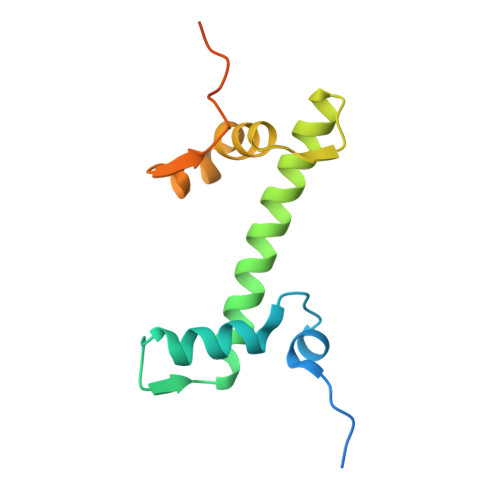
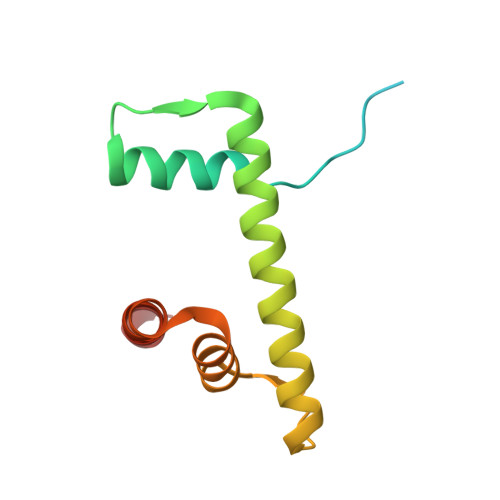
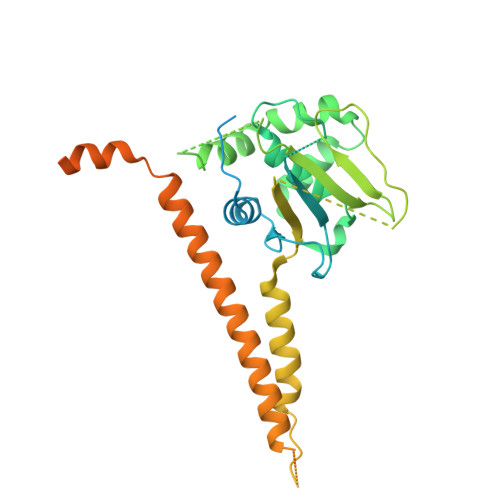
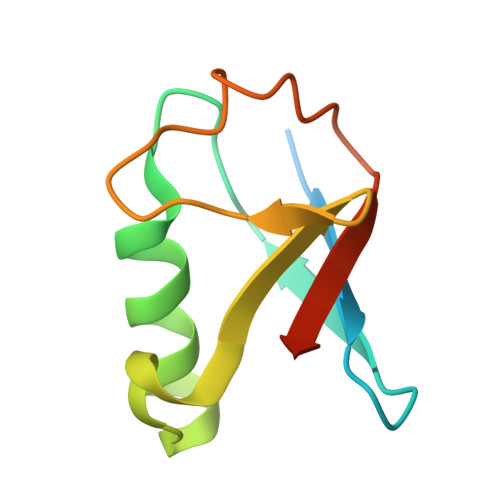
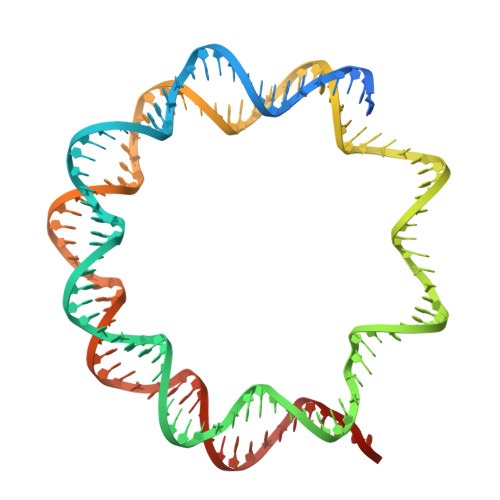
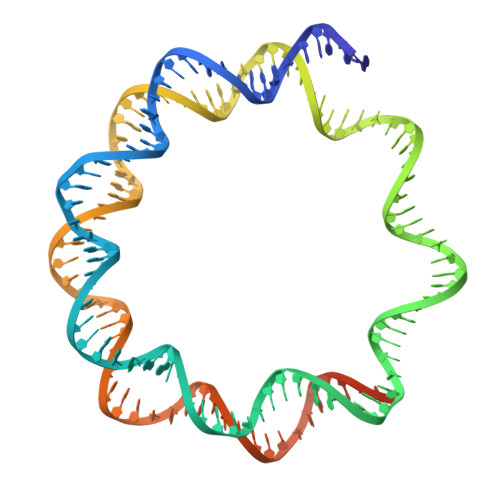
Structural basis of nucleosome deubiquitination by the bidentate Calypso/Asx complex.
Wang, C., Sun, F., Zhao, H., Zhang, N., Guan, J., Zhou, Y., Shuai, W., Zheng, H., He, J.(2026) iScience 29: 114958-114958
- PubMed: 41782825 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.isci.2026.114958
- Primary Citation Related Structures:
9V33 - PubMed Abstract:
The Polycomb repressive complex 1 (PRC1) and PR-DUB constitute a canonical pair of histone-modifying enzymes that deposit and remove monoubiquitinated H2A at lysine 119 (H2AK119ub1), serving as a model of dynamic epigenetic regulation. In humans, PR-DUB, composed of BAP1 and ASXL1, functions as a monomeric complex, while the Drosophila homolog Calypso/Asx forms a bidentate dimer (Calypso 2 : Asx 2 ) with an unclear chromatin engagement mechanism. Here, we present its cryo-EM structure bound to a nucleosome, revealing the molecular basis of interaction. Surprisingly, only one Calypso/Asx unit engages the nucleosome in a conformation similar to human BAP1/ASXL1, while the second remains disengaged. Structural and biochemical analysis of the positively charged Calypso C terminus suggests a "spreading" potential of the bidentate complex along chromatin, which was validated in vitro using nucleosome arrays. These findings support a model in which the bidentate Calypso/Asx complex enables processive deubiquitination along chromatin via alternating or cooperative engagement.
- Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei, Anhui 230027, China.
Organizational Affiliation: