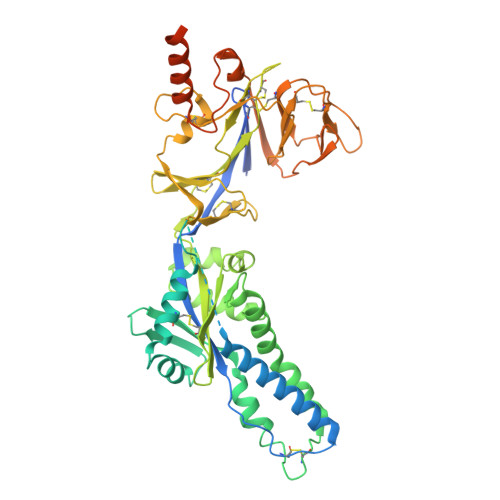
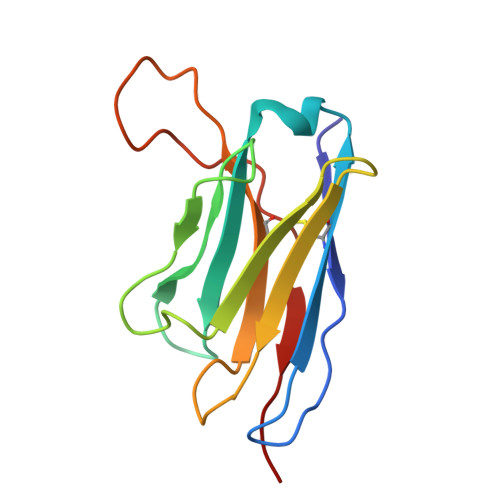
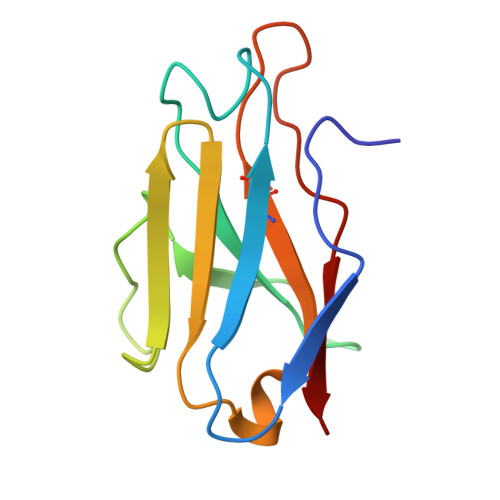
Antibody cocktails based on the occupationally acquired immunity of pediatricians neutralize and confer protection against RSV and hMPV.
Zhai, H., Yu, W., Wang, J., Deng, J., Lei, S., Zhou, T., Li, Y., Xu, K., Ma, M., Feng, R., Hu, Y., Ren, L., Cao, Y., Liu, E., Wang, X.(2026) Sci Transl Med 18: eadz4170-eadz4170
- PubMed: 41706868 Search on PubMed
- DOI: https://doi.org/10.1126/scitranslmed.adz4170
- Primary Citation Related Structures:
9V0N, 9V0P, 9V0Q, 9V0S, 9V0T, 9V2O, 9V2Q, 9V2R - PubMed Abstract:
Human respiratory syncytial virus (RSV) and human metapneumovirus (hMPV) are major causes of severe respiratory infections in young children, older adults, and immunocompromised individuals. Here, we isolated RSV fusion (F) protein-specific B cells from pediatricians who are routinely exposed to these viruses. We then derived monoclonal antibodies (mAbs) from those B cells to characterize their binding and neutralization profiles. Among the isolated mAbs, we found that CNR2056 and CNR2053 (targeting site Ø of the pre-F protein) potently neutralized diverse RSV A and B strains; another mAb, CNR2047 (targeting site III), uniquely exhibited cross-neutralization capacity against both RSV and hMPV variants. In vivo, prophylactic administration of CNR2056 and CNR2053 controlled lung viral loads and pathology in RSV A2- and B9320-challenged cotton rats. Moreover, a prophylactic dose of 0.5 milligrams per kilogram of CNR2047 resulted in complete protection against hMPV in the lungs of BALB/c mice. Structural analysis revealed unique binding modes for the three mAbs, supporting the potential for rational mAb cocktail design. Deep mutational scanning for RSV F further demonstrated that mutations required to evade CNR2053 and CNR2056 were primarily in evolutionarily constrained sites, suggesting a fitness cost to immune escape. Rationally combining site Ø- and site III-directed mAbs (e.g., CNR2056-CNR2047) into cocktails conferred additive effects, expanding coverage to hMPV and minimizing risk of escape variants. Thus, rationally designed cocktails of CNR2056, CNR2053, and CNR2047 may offer a versatile immunoprophylactic agent against a range of pneumoviruses with potential to protect against both current and future variants.
- Department of Respiratory Medicine, Children's Hospital of Chongqing Medical University, National Clinical Research Center for Children and Adolescents' Health and Diseases, Ministry of Education Key Laboratory of Child Development and Disorders, Chongqing 400014, China.
Organizational Affiliation: