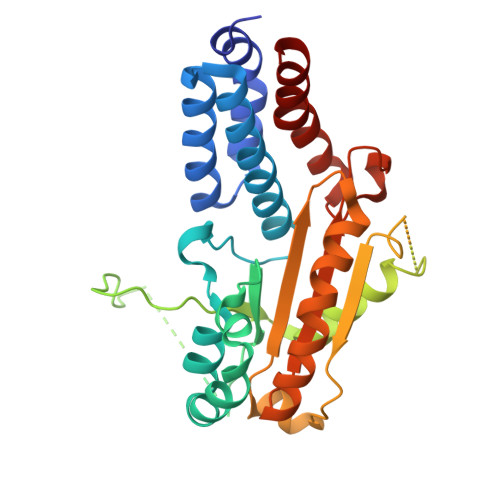
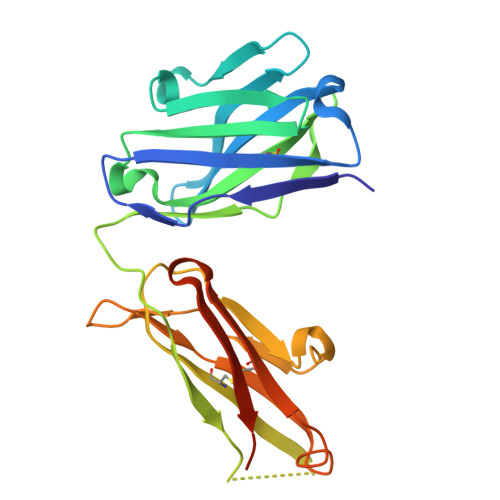
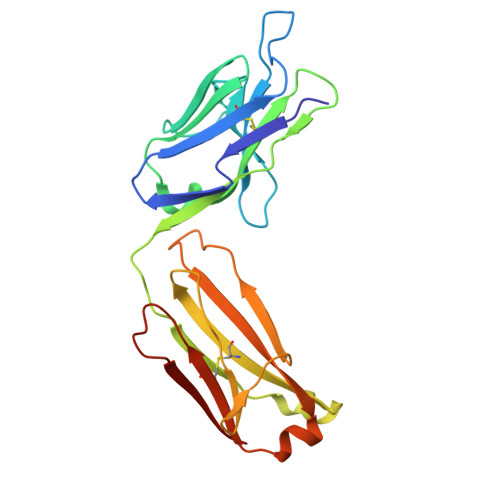
Structure and function of the nairovirus cap-snatching endonuclease.
Kuang, W., Tian, Z., Zhang, G., Wu, F., Li, J., Tang, J., Zhang, H., Zhuo, X., Hu, Z., Wang, M., Zhao, H., Deng, Z.(2026) Nucleic Acids Res 54
- PubMed: 41533578 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaf1515
- Primary Citation Related Structures:
9UZA, 9UZB, 9UZC, 9UZD, 9UZE, 9UZF, 9UZG, 9UZH, 9UZI - PubMed Abstract:
Nairoviruses include several human pathogens such as Crimean-Congo hemorrhagic fever virus (CCHFV) and Kasokero virus (KASV). The cap-snatching endonuclease (EN) domain of the viral polymerase is essential for transcription and represents a promising antiviral target. However, the structural and functional mechanisms of nairovirus ENs remain poorly understood. Here, we describe biochemical and structural studies of the ENs from CCHFV and KASV. Biochemical assays demonstrate that the RNA endonuclease activity of both ENs is activated by manganese ions and exhibits a preference for uridine-rich RNA substrates. This activity is inhibited by three metal-chelating inhibitors (DPBA, L-742,001, and BXA), with BXA displaying the highest binding affinity and inhibitory potency. We further determine nine crystal structures of CCHFV and KASV ENs in apo, metal ion-bound, and inhibitor-bound states. Comparative structural analysis uncovers a two-metal-ion binding mode unique to nairovirus ENs, in which conserved residues coordinate two manganese ions via bridging water molecules. In the inhibitor-bound structures of KASV EN, BXA forms additional stabilizing interactions with the enzyme, explaining its superior inhibitory effect. Functional assays further confirm that the two-metal-ion mechanism is critical for viral transcription. These findings provide a structural foundation for the rational design of antivirals against CCHFV and related pathogens.
- State Key Laboratory of Virology and Biosafety, Wuhan Institute of Virology, Chinese Academy of Sciences, Wuhan 430207, Hubei, China.
Organizational Affiliation: