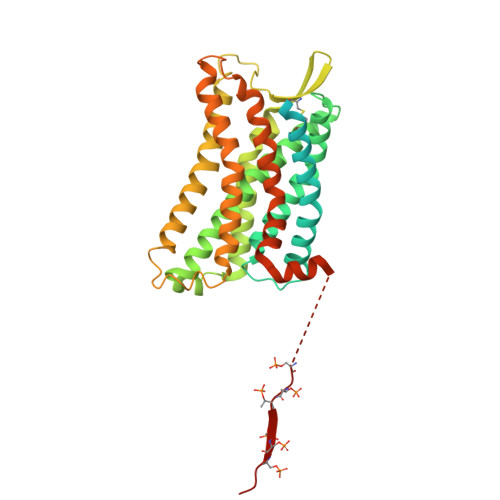
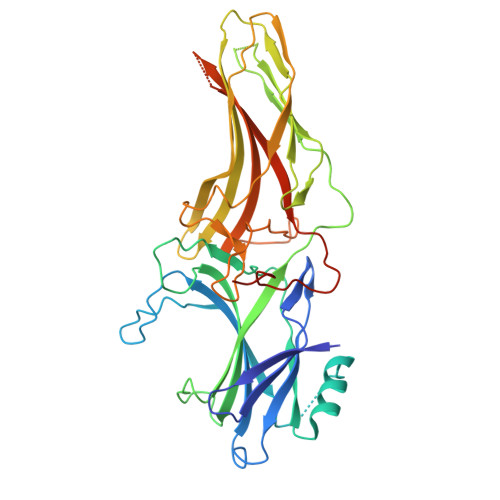
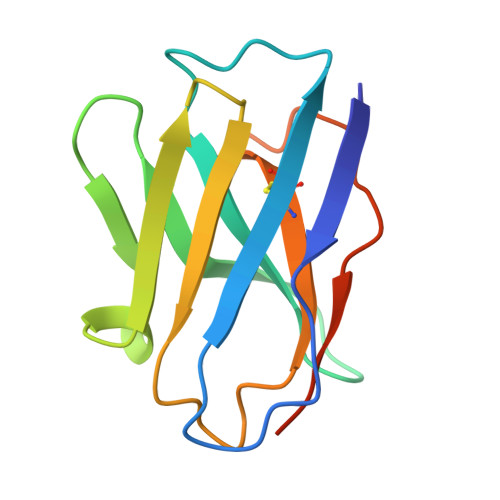
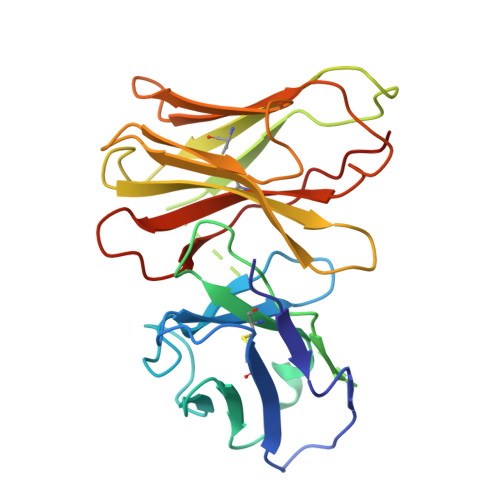
Noncanonical agonist-dependent and -independent arrestin recruitment of GPR1.
Cai, H., Lin, X., Zhao, L., He, M., Yu, J., Zhang, B., Ma, Y., Chang, X., Tang, Y., Luo, T., Jiang, J., Ma, M., Song, W., Ma, L., Chu, X., Yi, C., Chen, K., Han, S., Xie, C., Shui, W., Zhao, Q., Zhu, Y., Wu, B.(2025) Science 390: eadt8794-eadt8794
- PubMed: 41264711 Search on PubMed
- DOI: https://doi.org/10.1126/science.adt8794
- Primary Citation Related Structures:
9UYH, 9UYI, 9UYJ, 9UYL, 9UYM, 9UYN - PubMed Abstract:
G protein (heterotrimeric guanine nucleotide-binding protein)-coupled receptors have diverse signaling properties with differential preferences for downstream pathways. Certain receptors, such as the chemerin receptor GPR1, undergo arrestin-mediated internalization but weak G protein signaling. However, the mechanisms of this unusual signaling pattern and its physiological relevance are unclear. We report the structures of GPR1 bound to chemerin and β-arrestin 1 or β-arrestin 2 and an agonist-free GPR1-β-arrestin 1 complex. Upon agonist stimulation, the receptor binds the two arrestins in distinct interaction patterns, which may account for their differential cellular responses. Agonist-independent internalization was mediated by an inactive, constitutively phosphorylated GPR1 that accommodates β-arrestin 1 in an unconventional pocket together with a fatty acid, which potentially provides a basis for GPR1 modulating lipid accumulation in lipid-overloaded adipocytes.
- School of Pharmaceutical Science and Technology, Hangzhou Institute for Advanced Study, University of Chinese Academy of Sciences, Hangzhou, China.
Organizational Affiliation: