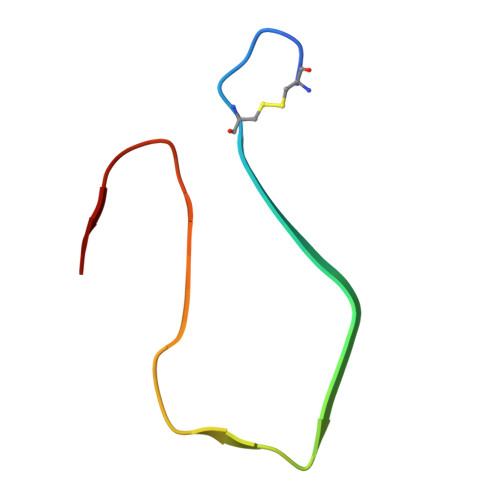
Structure of pancreatic hIAPP fibrils derived from patients with type 2 diabetes.
Liu, W., Han, J., Gong, W., Zhang, F., Cao, Q.(2026) Cell 189: 1201
- PubMed: 41483806 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2025.12.001
- Primary Citation Related Structures:
9ULZ - PubMed Abstract:
Type 2 diabetes (T2D) impacts the quality of life and lifespan of nearly 10% of the global population. Human islet amyloid polypeptide (hIAPP) constitutes a major component of islet amyloid deposition in patients with T2D, with hIAPP fibrils believed to play a key role in the pathogenesis of T2D. In this study, we determined the cryo-electron microscopy (cryo-EM) structure of hIAPP fibrils extracted from surgically resected pancreases of three donors with T2D. These fibrils exhibit a uniform morphology, comprising two symmetrical protofilaments encompassing residues 2-37 of hIAPP and adopting an Ω-shaped fold. The structure of pancreatic hIAPP fibrils differs from that of fibrils formed in vitro. Additional densities were observed in the pancreatic hIAPP fibrils, suggesting ligand binding that may play significant roles in the pathogenesis of T2D. Collectively, our study presents the atomic structure of pathological hIAPP fibrils, contributing to the therapeutic and mechanistic exploration of T2D.
- Bio-X Institutes, Key Laboratory for the Genetics of Developmental and Neuropsychiatric Disorders, Ministry of Education, Shanghai Jiao Tong University, Shanghai 200030, China.
Organizational Affiliation: