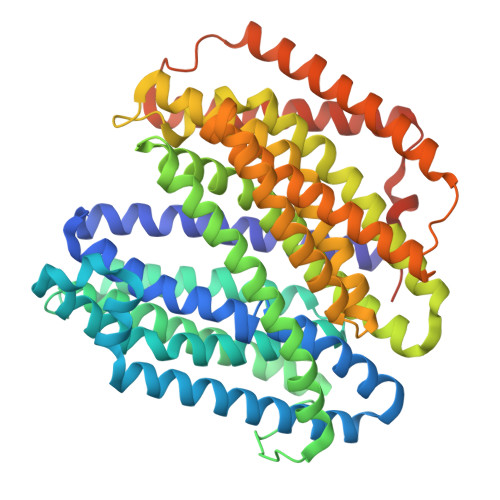
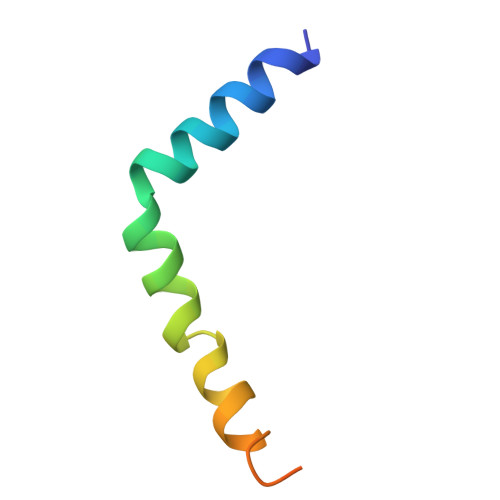
Phage lysis protein Lys M acts as a wedge to block MurJ conformational changes.
Kohga, H., Lertpreedakorn, N., Miyazaki, R., Wu, S., Hosoda, K., Tanaka, H., Takahashi, Y.S., Yoshikaie, K., Kuruma, Y., Shigematsu, H., Mori, T., Tsukazaki, T.(2025) Sci Adv 11: eady8083-eady8083
- PubMed: 41061077 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.ady8083
- Primary Citation Related Structures:
9UKV - PubMed Abstract:
Many antibiotics target essential cellular processes. To combat multidrug-resistant bacteria, new antibacterial strategies are needed. In the peptidoglycan biogenesis pathway in Escherichia coli , MurJ, the lipid II flippase, is an essential membrane protein. The 37-residue protein M from the Levivirus phage, known as Lys M or Sgl M , targets MurJ and induces cell lysis; however, its molecular mechanism remains unclear. Here, we present the cryo-EM structure of the MurJ/Lys M (JM) complex at 3.09-angstrom resolution, revealing that Lys M interacts with the crevasse between TM2 and TM7 of MurJ, locking MurJ in an outward-facing conformation, with Lys M acting like a wedge. Alanine-scanning mutagenesis and pull-down assays revealed key residues responsible for Lys M function, and molecular dynamics simulations showed that Lys M stabilizes MurJ's outward-facing state. These findings demonstrate an unprecedented phage-derived mechanism for blocking lipid II transport, providing a structural framework for designing MurJ-targeted antimicrobial agents.
- Nara Institute of Science and Technology, Ikoma, Nara, Japan.
Organizational Affiliation: