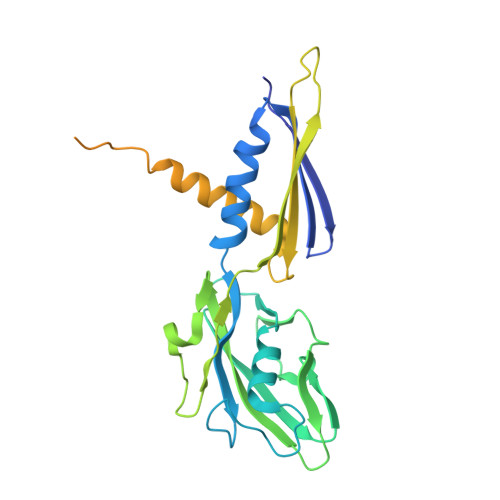
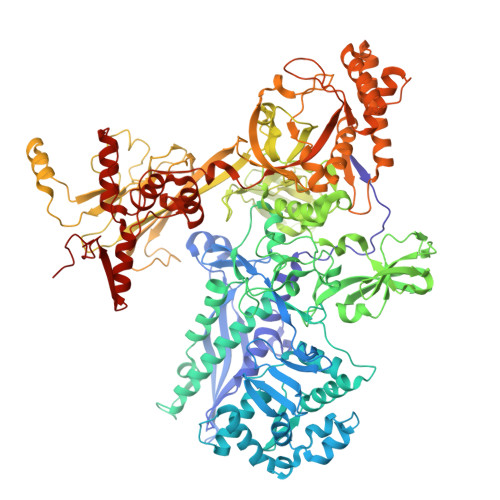
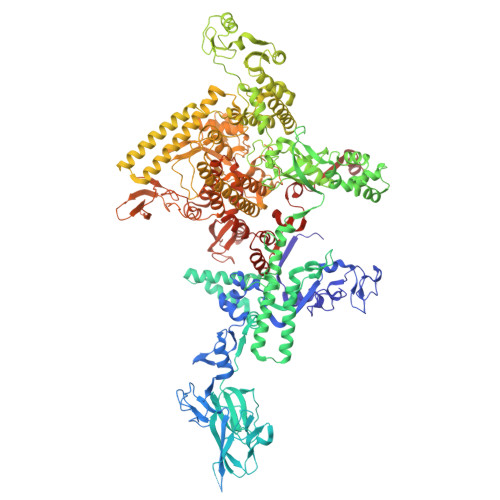
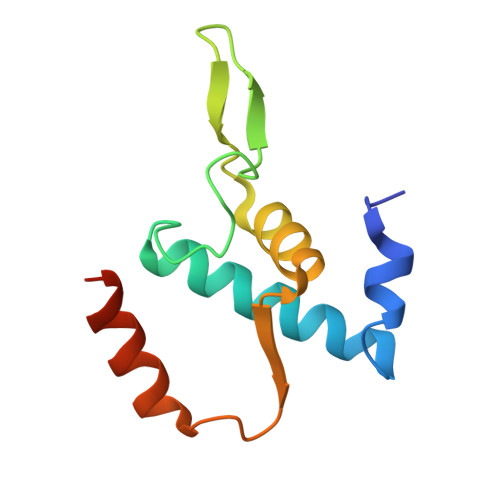
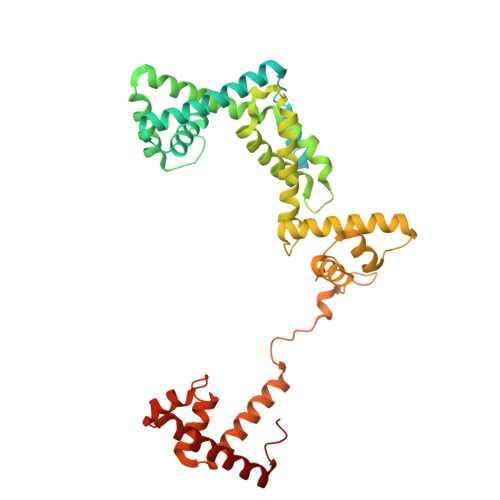
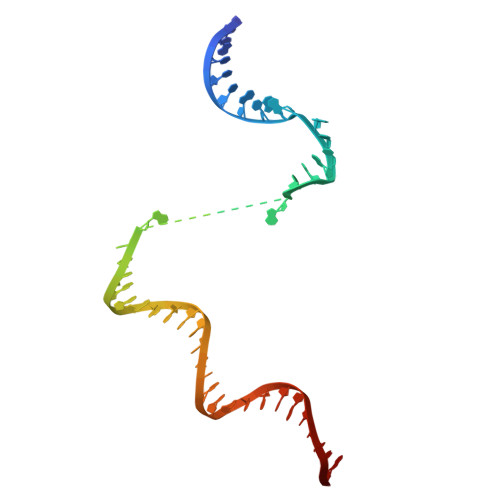
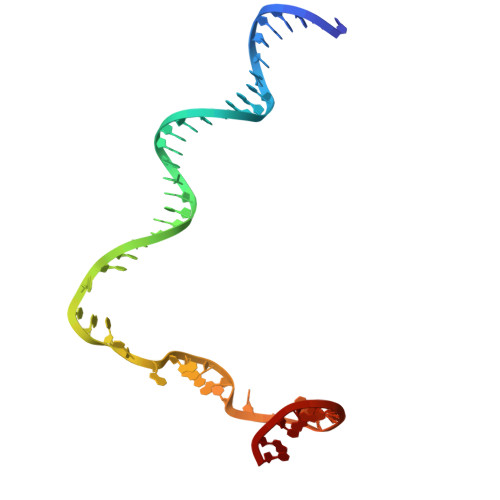
Molecular basis for noncanonical transcription initiation from Np 4 A alarmones.
Duan, W., Kaushik, A., Unarta, I.C., Wu, Y., Liu, M.M.J., Weaver, J.W., Wang, B., Rice, W.J., Luciano, D.J., Belasco, J.G., Huang, X., Nudler, E., Serganov, A.(2025) Nat Chem Biol
- PubMed: 41094128 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-025-02044-6
- Primary Citation Related Structures:
9UJK, 9UJL, 9UJN, 9UJP, 9UKN, 9UKO, 9UKP, 9UKS, 9UKT, 9UKU, 9ULS, 9ULT, 9UMA, 9UPW, 9V4M - PubMed Abstract:
Stress-induced dinucleoside tetraphosphates (Np 4 Ns, where N is adenosine, guanosine, cytosine or uridine) are ubiquitous in living organisms, yet their function has been largely elusive for over 50 years. Recent studies have revealed that RNA polymerase can influence the cellular lifetime of transcripts by incorporating these alarmones into RNA as 5'-terminal caps. Here we present structural and biochemical data that reveal the molecular basis of noncanonical transcription initiation from Np 4 As by Escherichia coli and Thermus thermophilus RNA polymerases. Our results show the influence of the first two nucleotide incorporation steps on capping efficiency and the different interactions of Np 4 As with transcription initiation complexes. These data provide critical insights into the substrate selectivity that dictates levels of Np 4 capping in bacterial cells.
- Department of Biochemistry and Molecular Pharmacology, New York University School of Medicine, New York, NY, USA.
Organizational Affiliation: