Structure of HN HA at pH 7.0 (conformation2).
Deng, G.J., Sun, H.L., Peng, Q.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Hemagglutinin | A, C [auth B], E [auth C] | 327 | Influenza A virus | Mutation(s): 0 Gene Names: HA | 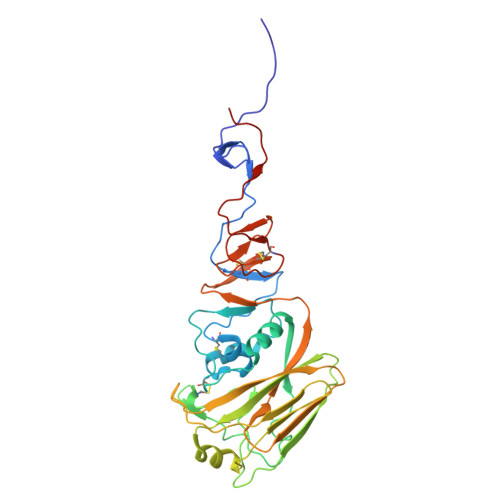 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A6G5UYK1 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Hemagglutinin | B [auth D], D [auth E], F | 222 | Influenza A virus | Mutation(s): 0 Gene Names: HA | 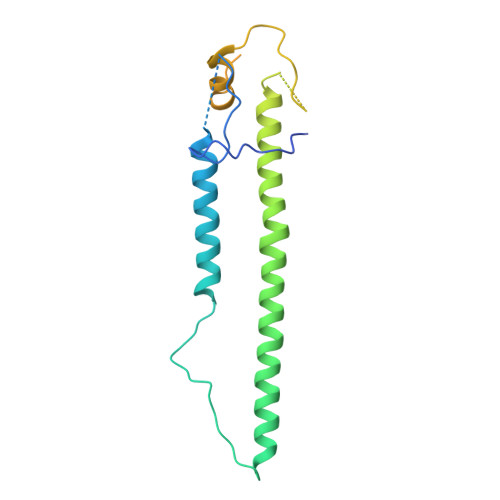 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0A6B7KK93 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Task | Software Package | Version |
|---|---|---|
| RECONSTRUCTION | cryoSPARC |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | 32200752 |