Crystal structure of a phenylalanine-modifi ed thrombin-binding DNA aptamer
Park, S., Kanazawa, H., Kondo, J.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Find similar nucleic acids by: Sequence
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | Organism | Image | |
| DNA (5'-D(*GP*G*(FNH)P*TP*GP*GP*TP*GP*TP*GP*GP*TP*TP*GP*G)-3') | A [auth B] | 15 | synthetic construct | 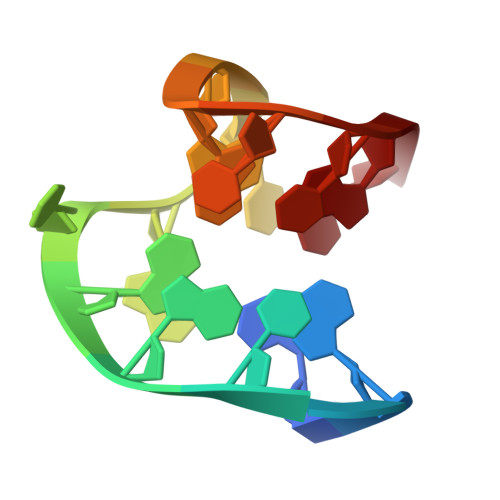 | |
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PHE (Subject of Investigation/LOI) Query on PHE | B | PHENYLALANINE C9 H11 N O2 COLNVLDHVKWLRT-QMMMGPOBSA-N |  | ||
| BA Query on BA | C [auth B] | BARIUM ION Ba XDFCIPNJCBUZJN-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 32.055 | α = 90 |
| b = 54.44 | β = 90 |
| c = 21.419 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XSCALE | data scaling |
| XDS | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Japan Agency for Medical Research and Development (AMED) | Japan | -- |