The low-density lipoprotein receptor LDLR mediates cellular entry of nonenveloped hepatitis A virus.
Shiota, T., Zhao, Y., Shiota, I., Duyvesteyn, H.M.E., Yamaoka, M., Das, A., Kapustina, M., Shah, P., Muramatsu, M., Wang, X., Fry, E.E., Stuart, D.I., Lemon, S.M.(2026) Proc Natl Acad Sci U S A 123: e2534261123-e2534261123
- PubMed: 41894341 Search on PubMed
- DOI: https://doi.org/10.1073/pnas.2534261123
- Primary Citation Related Structures:
9TBH, 9TBI - PubMed Abstract:
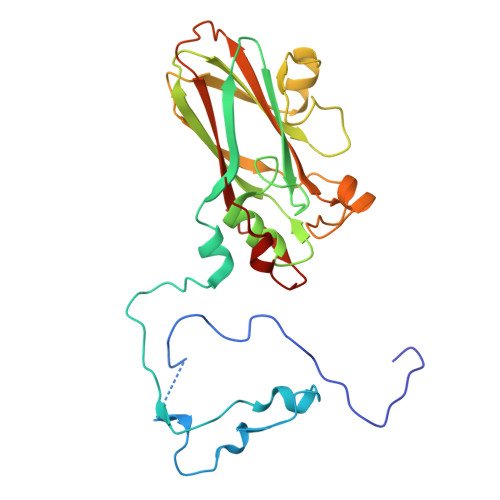
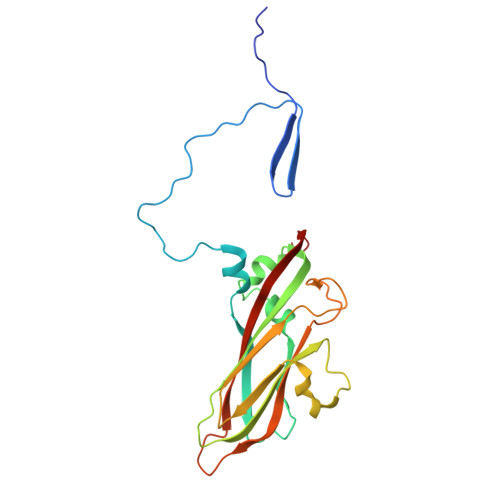
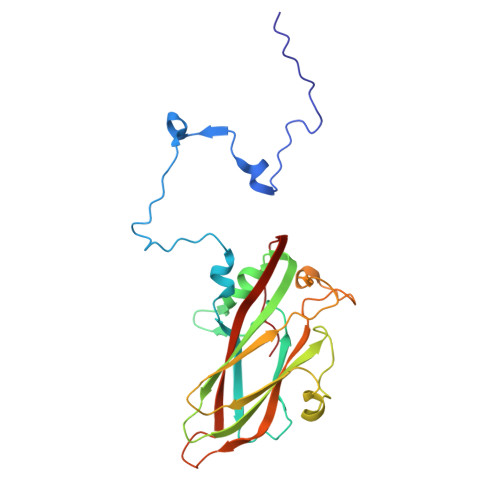
Hepatitis A virus (HAV) is an unusual picornavirus with two types of extracellular virions: nonenveloped particles (nHAV) shed in feces and quasi-enveloped particles (eHAV) circulating in blood. Both enter cells by clathrin-dependent endocytic pathways merging in late endolysosomes with capsid binding to ganglioside receptors. Phosphatidylserine receptors facilitate eHAV endocytosis, but no protein receptor has been identified for nHAV. Here, we show low-density lipoprotein receptor (LDLR) is such a receptor. LDLR knockout did not alter viral attachment to cells, but restricted cellular uptake of nHAV (not eHAV). Soluble LDLR ectodomain blocked nHAV entry, as did antibody to LDLR. Recombinant LDLR-related protein-associated protein 1, a pan-LDLR family chaperone, also inhibited nHAV entry, including residual entry into knockout cells, suggesting other LDLR family members may similarly facilitate endocytosis. Reconstituting full-length LDLR expression restored nHAV entry in knockout cells, whereas LDLR mutants lacking LA repeats 4 to 7 or the EGF-like/propeller domain did not. ELISAs confirmed LDLR binds nHAV, optimally above pH7, without destabilizing the capsid. A 1.7Å resolution cryoelectron microscopy (cryo-EM) structure revealed LDLR interacts with VP1 at the fivefold vertex of the capsid. Extreme blurring of the LDLR density prevented detailed identification of LDLR interactions, and suggested binding does not follow particle symmetry, being either flexible or to multiple LDLR regions. Additional cryo-EM studies show ganglioside GD1a binds to a similar region of the capsid. Collectively, these data reveal the LDLR to be an important entry factor, shuttling nHAV from the extracellular environment to endolysosomes where it is likely released at low pH to bind gangliosides.
- Lineberger Comprehensive Cancer Center, The University of North Carolina at Chapel Hill, Chapel Hill, NC 27599.
Organizational Affiliation: